the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Reconstruction of δ13CDIC in the Atlantic Ocean: a probabilistic machine learning approach for filling historical data gaps
Hui Gao
Zhentao Sun
Diana Cai
Meibing Jin
Stable carbon isotope composition of marine dissolved inorganic carbon (DIC), δ13CDIC, is a valuable tracer for oceanic carbon cycling. However, its observational coverage remains much sparser than that of DIC and other physical or biogeochemical variables, limiting its full potential. Here, we reconstruct δ13CDIC in the Atlantic Ocean using a probabilistic machine learning framework, Gaussian Process Regression (GPR). We compiled data from 51 historical cruises, including a high-resolution 2023 A16N transect, and applied secondary quality control via crossover analysis, retaining 37 cruises for model training, validation, and testing. The trained GPR model achieved an average bias of −0.007 ± 0.082 ‰ and an overall uncertainty of 0.11 ‰, arising from measurement (0.07 ‰), mapping (0.08 ‰), and input-variable (0.009 ‰) errors. To address validation limitations related to sparse observations, we further supplemented this work with numerical model-based validation (Claret et al., 2021), confirming the GPR model's robustness in δ13CDIC reconstruction. Using the GLODAPv2.2023 Atlantic dataset as predictors, the reconstruction expanded the number of acceptable δ13CDIC samples by a factor of 7.65, from 8941 to 68 435 across the Atlantic basins. The resulting dataset markedly improves the spatial resolution in longitude, latitude, and depth, and provides enhanced temporal continuity over the past four decades, offering great advantages in decadal trend assessment. Compared to the sparse original measurements, the reconstruction also reduces spatial discontinuities and reveals finer vertical structures consistent with other high-resolution biogeochemical observations. Additionally, the validated GPR framework was applied to the GLODAPv2 1° × 1° global interior ocean mapped climatology (Lauvset et al., 2016), producing a climatological gridded 3D δ13CDIC dataset for the Atlantic Ocean. These reconstructed δ13CDIC datasets provide new opportunities to resolve regional carbon cycle dynamics, validate Earth system models, refine estimates of oceanic carbon uptake on at least decadal timescales, and extend climate reanalysis records. The reconstructed δ13CDIC data, quality-controlled observational data from 51 cruises, and gridded δ13CDIC product are available at https://doi.org/10.5281/zenodo.18481145 (Gao et al., 2025).
- Article
(10901 KB) - Full-text XML
- BibTeX
- EndNote
The stable carbon isotope ratio has been widely applied as a tracer in marine carbon research, providing valuable insights into various processes within the oceanic carbon system. It is expressed via the standard delta notation: δ13C = ((13C 12C)C 12C, referenced against the international standard, the Vienna Pee Dee Belemnite (V-PDB) fossil. Specifically, the δ13C of dissolved inorganic carbon (DIC), denoted as δ13CDIC (expressed in per mil, ‰), has proven instrumental in studies encompassing estimating rates of biological production in surface ocean mixed layer (Quay et al., 2003, 2009; Yang et al., 2019), quantifying anthropogenic carbon inputs and accumulations in ocean basins (Quay et al., 1992, 2003, 2007, 2017; Körtzinger et al., 2003; Olsen and Ninnemann, 2010; Racapé et al., 2013), and validating earth system models (Sonnerup and Quay, 2012; Schmittner et al., 2013; Liu et al., 2021; Claret et al., 2021), making it an indispensable parameter in understanding the complexities of the marine carbon cycle.
Measurements of δ13CDIC in the ocean trace their roots to the mid-20th century, with significant advancements occurring in the 1970s and 1980s due to the development of more precise mass spectrometry techniques. A pivotal moment in marine isotope research came with Kroopnick's comprehensive analyses of δ13CDIC distribution in the Atlantic Ocean (Kroopnick, 1980a) and globally (Kroopnick, 1985), which provided critical insights into isotopic patterns across the oceans. Over subsequent decades, observational δ13CDIC data and their biogeochemical interpretation continue to increase (Gruber et al., 1999; Quay et al., 2003, 2017; Schmittner et al., 2013). The creation of databases such as Global Ocean Data Analysis Project (GLODAP) further enhanced access to δ13CDIC data and other carbon-related parameters (Olsen et al., 2016). However, unlike other carbon datasets such as DIC and total alkalinity (TA), δ13CDIC lacked secondary quality control until Becker et al. (2016) introduced an internally consistent δ13CDIC dataset for the North Atlantic, marking a significant step in addressing biases and improving data reliability. But still, unlike other carbon datasets, δ13CDIC lacks necessary sufficiently high spatial resolution for it to be effective in ocean carbon cycle research and as a tracer for anthropogenic carbon accumulation in the ocean.
Traditionally, δ13CDIC data have been acquired by preserving and transporting seawater samples to shore-based laboratories for analysis via Isotope Ratio Mass Spectrometry (IRMS). Although the IRMS approach is highly precise and accurate, it is labor-intensive and unsuitable for at-sea analysis, and incapable of measuring precise DIC concentrations concurrently. These limitations have significantly restricted the collection of δ13CDIC data, impeding efforts to characterize spatiotemporal variabilities and long-term trends of δ13CDIC. For example, in the Atlantic Ocean, only 6820 δ13CDIC measurements were collected across 32 cruises over 40 years, averaging just 213 samples per cruise (Becker et al., 2016). Specifically, along the A16N transect, approximately 500 samples were collected during the 1993 and 2013 cruises and only 38 surface water samples were collected during the 2003 cruise for land-based δ13CDIC analysis, showing a stark contrast to the fact that about 3000 DIC samples were analysed at sea per cruise. To overcome this severe bottleneck, the Cai Lab has developed a precise, rapid, and sea-going suitable δ13CDIC analytical method with a precision of better than ±0.05 ‰ based on the Cavity Ring-Down Spectroscopy (CRDS) stable carbon isotope analyzer, G2131-i (Su et al., 2019; Deng et al., 2022; Sun et al., 2024, 2025). This method has been extensively tested during several recent observations and studies. During a long (58 d) ocean cruise along A16N in 2023, approximately 3500 δ13CDIC samples were collected, with ∼ 3000 samples analysed at sea, alongside DIC observations. This progress has significantly surpassed the historical A16N datasets (Sun et al., 2025).
Despite extensive field data collection efforts, observations of δ13CDIC remain sparse compared to other inorganic carbon chemistry variables (e.g., DIC and TA). The distribution and variations of inorganic carbon chemistry within water masses are governed by both the source-water properties and physical and biogeochemical processes, resulting in region-specific stoichiometric relationships among inorganic carbon variables. These relationships are typically nonlinear and exhibit spatial and temporal variability, making it challenging to determine general distribution patterns. In recent decades, advancements in machine learning techniques coupled with accumulated observational data have facilitated numerous studies that interpolate inorganic carbon chemistry variables, particularly the partial pressure of CO2 (pCO2) due to its relatively high spatial coverage, from satellite data and reanalysis products (e.g., Landschützer et al., 2016; Gregor and Gruber, 2021; Roobært et al., 2024; Wu et al., 2025). These methodological developments present a promising opportunity to investigate the potential for deriving δ13CDIC data from more abundantly measured variables.
Given the limited and fragmented δ13CDIC dataset compared to other parameters such as DIC, and to fully utilize the δ13CDIC tracer approach for quantifying anthropogenic CO2 uptake by the ocean (Quay et al., 2017), the rate of ocean biological production (Esposito et al., 2019; Quay et al., 2009, 2020; Quay, 2023), and carbon cycling across the land-ocean interface (Alling et al., 2012; Kwon et al., 2021; Samanta et al., 2015), this study aims to reconstruct a high-resolution δ13CDIC dataset for the Atlantic Ocean using Gaussian Process Regression (GPR), a probabilistic machine-learning approach capable of capturing nonlinear relationships and spatial-temporal variability. This reconstruction integrates historical δ13CDIC observations in the Atlantic Ocean with new high-resolution data collected along the A16N transect in 2023 (Sun et al., 2025), the GLODAPv2 1°×1° global interior ocean mapped climatology (Lauvset et al., 2016), and the product of GLODAPv2.2023 (Lauvset et al., 2024). The final product consists of three components with comprehensive uncertainty analysis: (1) a quality-controlled δ13CDIC observational dataset compiled from 51 cruises with a crossover analysis using standardized protocols, and (2) a machine learning-reconstructed δ13CDIC dataset derived from other inorganic carbon chemistry variables, (3) a climatological 1°×1° gridded three-dimensional δ13CDIC dataset for the Atlantic Ocean produced by applying the validated GPR framework to the GLODAPv2 1°×1° global interior ocean mapped climatology. The structure of this study is as follows: Sect. 2 describes the datasets used, the secondary quality control of δ13CDIC, and the methodology for reconstructing the δ13CDIC dataset. Section 3 evaluates the accuracy, performance, and applicability of the reconstructed δ13CDIC dataset in resolving its spatial and temporal distribution. Section 4 presents the conclusions. Sections 5 and 6 provide access the dataset, the codes used for its generation, and the figures presented in this study. The Appendix provides additional evaluation and validation by applying the same GPR approach to numerical model outputs.
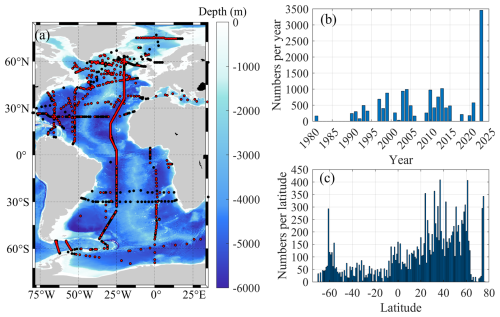
Figure 1Overview of the collected δ13CDIC data. (a) Map of all stations with δ13CDIC data in the Atlantic Ocean, with stations containing samples deeper than 2000 m highlighted in red. (b) Temporal distribution of δ13CDIC data, organized by year of collection. (c) Total number of δ13CDIC samples aggregated by each degree of latitude.
2.1 Data collection
2.1.1 Historical δ13CDIC Data Collection
In the Atlantic Ocean, δ13CDIC data were compiled from several international research initiatives, including GLODAP, Ocean Carbon and Acidification Data System (OCADS), Climate and Ocean Variability, Predictability, and Change (CLIVAR), Carbon Hydrographic Data Office (CCHDO) and the internally consistent dataset of δ13CDIC in the North Atlantic Ocean (NAC13v1, Becker et al., 2016). Notably, most cruises in the final compiled dataset are sourced from GLODAP. Cruises from the other aforementioned databases were only included to supplement those not covered by GLODAP, and no duplicate cruises were counted to ensure each unique cruise is represented exactly once. From the original dataset published by Becker et al. (2016), we excluded four cruises: 35TH20060521, 74JC20120601, 74DI20140606, and OMEX1NA, due to missing essential corresponding parameters, i.e., variables used for model training. The remaining cruises, including those from other sources and the 28 cruises retained from Becker et al. (2016), comprise a total of 51 cruises, covering 369 stations and 15 225 δ13CDIC samples (Fig. 1 and Table 1). To ensure internal consistency, samples from depths greater than 2000 m were selected for crossover analysis. Specifically, deep waters below 2000 m in the South Atlantic Ocean are most likely not impacted by anthropogenic carbon (i.e., Gao et al., 2022, 2024), supporting this threshold. In contrast, North Atlantic Deep Water (NADW) formation may drive small but measurable decadal anthropogenic carbon changes even below 2000 m. However, Becker et al. (2016) showed that restricting analysis to depths > 2000 m effectively reduces cruise offset variability in variable North Atlantic regions (e.g., Labrador Sea, Nordic Seas), further validating our 2000 m threshold for the Atlantic. This criterion resulted in 3772 samples from 305 stations deeper than 2000 m (highlighted as red points in Fig. 1a). The temporal and latitudinal distributions of the δ13CDIC data are illustrated in Fig. 1b and c, respectively. These datasets span from 1981 to 2023, with the most comprehensive annual δ13CDIC sampling occurring along A16N in 2023 (Fig. 1b). However, the sampling is spatially and temporally uneven, that is, data are sparse in certain years and latitudes, which underscores the need for an approach capable of generating robust predictions and uncertainty estimates in poorly sampled regions, such as the GPR method applied in this study. In particular, the majority of samples are concentrated in the North Atlantic, particularly between latitudes 25 and 60° N (Fig. 1c).
2.1.2 δ13CDIC Data Collection Along A16N in 2023
The A16N cruise in 2023 achieved extensive collection and high-resolution analysis of δ13CDIC and DIC datasets over two legs using onboard analytical techniques. Samples were collected from CTD rosette bottles into 250 mL borosilicate glass bottles following PICES best practices, preserved with HgCl2 to prevent biological activity, and sealed to prevent gas exchange. Analytical measurements were conducted onboard using two coupled systems comprising a CO2 extraction device (AS-D1 δ13CDIC Analyzer) and a CRDS (Picarro G2131-i), which simultaneously measured DIC concentrations and δ13CDIC values with high precision. Quality control measures included frequent calibration using Certified Reference Materials (CRMs) and in-house standards verified by IRMS, ensuring high data reliability with deviations mostly within ±0.03 ‰ for δ13CDIC (Sun et al., 2025).
2.2 Standardized Protocols for Crossover analysis
Secondary quality control of δ13CDIC data through crossover analysis ensures consistency across multi-cruise datasets (Becker et al., 2016; Tanhua et al., 2010; Lauvset and Tanhua, 2015). The analysis involves compiling data from overlapping regions, identifying crossover points within 222 km, and comparing δ13CDIC profiles in deep waters (> 2000 m) where variability is minimal. Profiles are interpolated to standard depths or density. In this study, we adopted the density-based interpolation: standard sigma4 surfaces (potential density referenced to 4000 m) are generated at 0.05-unit intervals, covering all observed densities, based on the interpolated density profile of the deepest station, following the workflows presented in Lauvset and Tanhua (2015). Mean offsets between overlapping profiles at the selected standard densities are calculated. Detailed workflows were. Systematic biases are identified using least squares minimization, and adjustments are proposed to align datasets without erasing real temporal or spatial trends. Adjustments are validated against known regional patterns and applied only if they reduce discrepancies to within the measurement uncertainty. All steps and corrections are documented to ensure transparency, resulting in reliable δ13CDIC datasets for carbon cycle analysis. Building on the NAC13v1 dataset provided by Becker et al. (2016), we propose additional adjustment recommendations for these cruises (Table 1). Given that cruise 64TR19900417 crossovers with cruise 33MW19930704, 06MT20040311, 33RO20130803, 33RO20230413, showing a very high mean offset and standard deviation −0.76 ± 0.23 ‰, and its δ13CDIC data is marked as NaN (missing values) in the NAC13v1 dataset, it is excluded from our analysis. After applying additional adjustments, the δ13CDIC data for the remaining 37 cruises exhibit high internal consistency. These 37 cruises do not include 13 cruises without deep-water crossover stations (marked as NC in the last column “Additional Adjustments” of Table 1) and cruise 64TR19900417, which were excluded to ensure data reliability as their uncertainties cannot be objectively quantified. Collectively, these excluded cruises accounted for less than 3 % of total δ13CDIC measurements. The internal accuracy of the adjusted δ13CDIC is determined to be −4.15 × 10−5 ‰ based on the calculation from Tanhua et al. (2010) and Becker et al. (2016). Finally, a total of 11 950 samples are used to train and test the model (Fig. 2). These samples were selected based on the quality flags of the relevant variables, where only data marked with quality flag values of 2 (acceptable measurement) or 6 (median of replicate measurements) were included, ensuring that the dataset is reliable and suitable for model development.
2.3 Model design
Predicting δ13CDIC in the ocean requires a method that can handle complex, highly nonlinear relationships and provide reliable uncertainty estimates for scientific interpretation. GPR (Seeger, 2004; Rasmussen and Williams, 2006) is particularly well suited to this task. As a non-parametric, probabilistic model, GPR not only produces point predictions but also quantifies uncertainty through credible intervals around the estimates. The versatility of GPR arises from its kernel function, which allows prior knowledge about the expected smoothness or variability of the δ13CDIC-environment relationship to be incorporated into the model, making GPR a powerful tool for robust predictions in oceanographic applications.
We employed GPR with a Matern kernel as the primary method for all subsequent δ13CDIC reconstructions, as it offers unique advantages tailored to addressing the core challenges of δ13CDIC data. The Matern class of kernels control function smoothness through its smoothness parameter. Specifically, the kernel yields functions that are smooth yet not overly restrictive, offering a balanced representation that aligns well with the expected variability of many data sets. Thus, it is expected to provide a balanced representation aligned with the physical variability of δ13CDIC in the ocean. Compared with the widely used squared exponential kernel, which assumes infinitely differentiable, and often unrealistically smooth functions, the Matern kernel allows for more plausible modeling of natural variability driven by ocean physical and biogeochemical processes such as mixing, air-sea CO2 exchange, and biological productivity. As a Bayesian non-parametric model, GPR also inherently balances model complexity and data fit to avoid overfitting to sparse, noisy observations, a critical benefit given the small sample sizes of many cruises and uneven spatial-temporal coverage in our dataset.
To evaluate this approach's performance, the Matern GPR was compared with a suite of alternative regression models, including GPR with other kernels, as well as additional baselines such as neural networks, support vector regression, and decision trees. A rigorous validation framework that balances robustness, feasibility, and adherence to synoptic cruise data characteristics was designed to underpin this comparison. Specifically, the dataset was randomly split into a training set (80 %) and a validation set (20 %), with model training and hyperparameter tuning performed using 10-fold cross-validation within the training set to mitigate overfitting. An independent test set was reserved for final performance evaluation, selected to ensure no overlap with the training/validation set in cruises, spatial regions, or temporal coverage. Our preference for random splitting (over cruise-separated k-fold cross-validation) initially stemmed from concerns that cruise-separated splitting would cause imbalanced folds, given the small sample sizes of many of the 37 cruises, which could lead to unstable hyperparameter tuning and biased results. Additionally, random splitting preserves the natural spatiotemporal variability of δ13CDIC, tuning the model to generalize across diverse oceanic conditions rather than overfitting to specific cruises. This core validation framework aligns with established practices for sparse oceanographic datasets (Lima et al., 2023; Regier et al., 2023; Wu et al., 2025).
To further enhance the statistical robustness of performance metrics and address the concern of synoptic cruise data characteristics, we additionally implemented a cruise-separated 5-fold cross-validation. To address the small sample sizes of some cruises, cruises were allocated to folds using an iterative balancing procedure that minimizes disparities in total sample size across folds to avoid biased results from imbalanced folds. Specifically, each cruise was assigned to the fold with the smallest current sample size, resulting in fold sample sizes ranging from 1836 to 2718 samples (6–8 cruises per fold). To eliminate data leakage, standardization was performed independently for each fold using only the fold's training subset, ensuring the validity of cross-validation results.
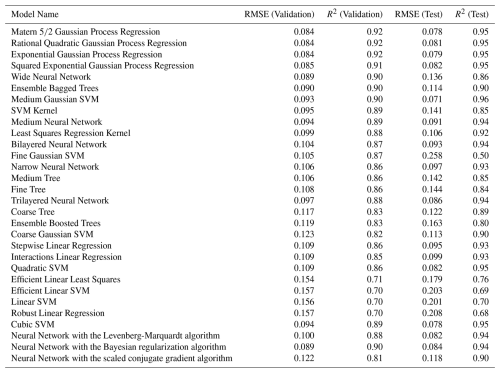
To clarify the division of labor between the two cross-validation strategies, the 10-fold cross-validation was retained exclusively for hyperparameter tuning, while the cruise-separated 5-fold cross-validation focused on evaluating robust performance across synoptic cruise units. Predictive performance for all models was assessed using the Root Mean Squared Error (RMSE) and the coefficient of determination (R2), computed separately for the validation and test sets. Among all tested models, including GPR with the squared exponential and other kernels (Table 2), the Matern GPR achieved the best predictive performance (lowest RMSE and highest R2) on both the validation set and the independent test set, while also providing meaningful uncertainty estimates, confirming its superiority for subsequent analysis.
Table 2Selection of machine-learning models based on Root Mean Squared Error (RMSE), and the coefficient of determination (R2).
The process of developing and reconstructing the δ13CDIC data product is outlined in Fig. 2. Data collection and preprocessing were conducted following the procedures detailed in Sect. 2.1 and 2.2.
The selected quality-controlled 37 cruises were used for model development, validation and test. Two of them (33RO20050111 and 33MW19930704), from the South and North Atlantic respectively, were randomly selected to form an independent test set (X1). The other 35 cruises formed dataset X2, which was further randomly split into a training set (80 %) and a validation set (20 %). The validation set was used to fine-tune hyperparameters and assess model performance, ensuring generalizability and helping identify overfitting (Wu et al., 2025). The model was trained using paired input variables: longitude (lon), latitude (lat), depth, temperature (T), salinity (S), apparent oxygen utilization (AOU), nitrate (N), silicate (Si), DIC, and atmospheric CO2 (xCO2), along with corresponding δ13CDIC values as the target variable (Eq. 1).
These input variables were selected to comprehensively capture the key physical, biological, and geochemical drivers influencing δ13CDIC. Specifically, longitude, latitude, and depth represent the spatial location of each observation, which is essential for resolving regional and vertical patterns. T and S reflect thermohaline forcing, i.e., the physical processes such as mixing, stratification, and water mass formation that impact carbon cycling. AOU, nitrate, and silicate are indicators of biological forcing, as they are influenced by biological productivity, remineralization, and nutrient utilization. DIC is directly related to δ13CDIC, since δ13CDIC reflects the stable carbon isotopic composition of the DIC pool; thus, variations in DIC are closely tied to changes in δ13CDIC. However, AOU, nutrients, and DIC also reflect partially physics as they are being mixed by ocean circulation. Finally, xCO2 represents external perturbations from air-sea CO2 exchange, which can alter both the concentration and isotopic composition of DIC. This term is necessary for predicting recent anthropogenic CO2 influences on ocean CO2 parameters, in particular, their decadal change rates.
The independent test set (X1), excluded entirely from training and validation, provides a final evaluation of model performance. Its role in assessing predictive skill across spatial data gaps ensures robust generalization. Finally, the trained GPR model is applied to the full set of hydrographic parameters from the GLODAPv2.2023 Atlantic dataset (https://glodap.info/index.php/merged-and-adjusted-data-product-v2-2023/, last access: 22 March 2026) to reconstruct a basin-wide δ13CDIC distribution over time across the Atlantic Ocean.
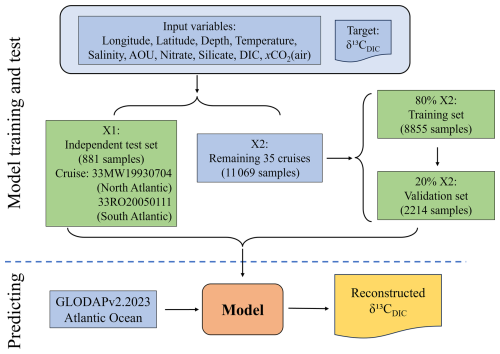
Figure 2A flowchart illustrating the machine-learning regression model for reconstructing the δ13CDIC product. The blue boxes represent the input, and the yellow box represents output datasets, while the green boxes depict the model training, validation, and independent testing processes. The orange box indicates the final trained model used for prediction.
2.4 Evaluation of model
The accuracy of the model outputs was evaluated using various statistical metrics, including the R2, RMSE, mean absolute error (MAE), and mean bias error (MBE). Among these metrics, MAE and MBE are valuable for evaluating the performance of the machine learning models. MAE quantifies the average absolute deviation between observed and predicted values; its insensitivity to outliers makes it ideal for handling the potential noise in δ13CDIC observational data, ensuring a robust measure of overall prediction error. MBE, by retaining the sign of deviations, identifies systematic biases (e.g., consistent overestimation or underestimation of δ13CDIC), which is critical for refining the machine learning model.
These metrics were calculated for the training, validation, and independent test phases, as defined below:
Here, i represents the ith sample, yobs,i refers to the observed δ13CDIC, is the mean of the observed δ13CDIC values, yest,i denotes the predicted δ13CDIC values from the final model, and N is the total number of matched samples.
2.5 Uncertainty of the reconstructed δ13CDIC
A comprehensive assessment of uncertainty of the reconstructed δ13CDIC was derived via error propagation, assuming independence between distinct uncertainty sources. These sources of uncertainties include: the direct δ13CDIC measurement uncertainty from observations (uobs), the uncertainty accumulated from the input variables (uinputs), and the uncertainty induced by the mapping function (umap). Following standard error propagation protocols (Hughes and Hase, 2010; Taylor, 1997), the comprehensive uncertainty of our estimated δ13CDIC product, , was calculated as the root sum of the squares of the individual uncertainties:
The observational uncertainty uobs inherent in δ13CDIC measurements varies by analytical method. For samples, when it is analyzed using IRMS, reported uncertainties range from ±0.12 ‰ (Gruber et al., 1999) to ±0.03 ‰ (Quay et al., 2003). For CRDS analysis, uncertainties are reported as ±0.07 ‰ for cruise 33RO20200321 (Gao et al., 2024) and ±0.03 ‰ for cruises 33RO20230306 and 33RO20230413 (Sun et al., 2025). Taking a conservative approach, we adopted an average uobs value of 0.07 ‰.
The input variable uncertainty (uinputs) accounts for uncertainties in temperature, salinity, nitrate, silicate, DIC, alkalinity, and xCO2. Monte Carlo simulation is performed 1000 times to quantify uinputs. Following Carter et al. (2024) and Wu et al. (2025), the perturbation of 0.002, 0.002, 2, 0.4, 0.4, 2 and 0.2 are randomly applied to temperature, salinity, AOU, nitrate, silicate, DIC, and xCO2, respectively. We then recalculated δ13CDIC using these perturbed inputs and quantified the resulting changes. The uncertainty contribution from each variable was determined as the standard deviation of the differences between the original reconstructed δ13CDIC and the noise-perturbed values. The final uinputs was calculated as the square root of the quadratic sum of these individual uncertainties.
The uncertainty associated with the mapping function, umap, is introduced within the GPR model. Here, umap is primarily quantified using the GPR model's posterior predictive standard deviation (i.e., the square root of the posterior predictive variance), that is, GPR uncertainty. GPR uncertainty is a conceptually direct measure that reflects the model's inherent uncertainty in mapping environmental predictors to δ13CDIC across the entire prediction set. The mean value of this GPR-derived umap is 0.08 ‰ across the prediction set, which captures both model variability and uncertainty related to generalization to unobserved regions. Wu et al. (2025) suggested that umap may alternatively be estimated as the RMSE between reconstructed and observed δ13CDIC on the reconstruction dataset, and we additionally report this RMSE as a reference to facilitate compatibility. Notably, the two metrics are quantitatively similar, ensuring our core uncertainty estimate remains consistent with widely adopted reporting practices while prioritizing the GPR-derived measure's scientific rigor.
3.1 Evaluation of the GPR model performance
The trained GPR model exhibited robust performance and high accuracy in representing δ13CDIC characteristics across the Atlantic Ocean (Fig. 3). During the training phase, we leveraged a 10-fold cross-validation approach, selected as it is a standard pre-implemented option in MATLAB's Machine Learning Toolbox. This approach balances computational efficiency and robustness, reducing overfitting by iteratively splitting training data into 10 folds: 9 for training and 1 for validation per iteration, with results averaged across iterations to ensure stable performance. Finally, the model with 10-fold cross-validation achieved an R2 of 0.92, an RMSE of 0.083 ‰, an MAE of 0.056 ‰, and an MBE of −0.0003 ‰ during training phase (Fig. 3a). Performance metrics during the validation phase (Fig. 3b) were comparable to those in the training phase, demonstrating the model's strong generalization ability, i.e., its capacity to accurately predict data not used for training, and confirming its resistance to overfitting. Additionally, in the independent testing phase, the model also maintained high accuracy (R2 = 0.95, RMSE = 0.082 ‰, MAE = 0.056 ‰, MBE = 0.007 ‰, Fig. 3c), with most samples clustered closely around the 1:1 line. These results indicate that the model can reliably predict δ13CDIC across diverse, unobserved spatial and temporal scales.
To further verify the model's robustness across different cruise combinations, we additionally conducted a cruise-separated 5-fold cross-validation. This supplementary cross-validation also yielded stable performance across all folds: per-fold RMSE ranged from 0.097 ‰ to 0.156 ‰ with an average of 0.124 ‰ (±0.027 ‰, standard deviation), and per-fold adjusted R2 ranged from 0.74 to 0.92 with an average of 0.80 (±0.08, standard deviation). The small standard deviations of these metrics indicate that the model's performance is not sensitive to the specific combination of training cruises, confirming no significant overfitting caused by uneven sample sizes across cruises and further reinforcing the reliability of the training results. Collectively, these findings, including stable performance in 10-fold cross-validation, reliable results from the cruise-separated 5-fold cross-validation, consistent accuracy in the validation phase, and high reliability in independent testing, demonstrate that the GPR-based δ13CDIC reconstruction method is highly generalizable and robust, enabling it to accurately capture the characteristics in δ13CDIC and provide reliable predictions across the Atlantic Ocean.
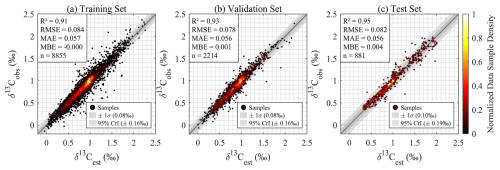
Figure 3Regression model evaluation for δ13CDIC reconstruction: Density scatter plots comparing model-estimated (δ13Cest) versus observed (δ13Cobs) δ13CDIC values during (a) training (80 % samples from 35 cruises, with 10-fold cross-validation), (b) validation (20 % samples from 35 cruises), and (c) independent testing (samples from cruises 33RO20050111 and 33MW19930704). Statistical metrics include coefficient of determination (R2), root-mean-square error (RMSE), mean absolute error (MAE), mean bias error (MBE), and sample size (n). Color indicates normalized local data point density within 2D bins: the plot area is divided into a 100 × 100 grid, raw density is the number of points per bin, and values are normalized to 0∼ 1 (0= sparsest, 1 = densest). Shaded bands represent predictive uncertainty, with the darker grey band showing one standard deviation (±1σ) and the lighter grey band showing the 95 % credible interval (CrI).
3.2 Evaluation of the spatial distribution of δ13CDIC
To assess the product's ability to capture δ13CDIC spatial patterns and quantify biases, we utilized the δ13CDIC distribution from independent test cruises 33MW19930704 and 33RO20050111 (Fig. 4). Both cruises are part of the repeated observation A16, specifically A16N (1993) and A16S (2005), which traverses the entire Atlantic Ocean from the sub-Arctic to the Southern Ocean (Fig. 4a). This transect has been frequently used in existing research and textbooks to represent Atlantic-scale distributions of carbonate chemistry and other characteristics (i.e., Wanninkhof et al., 2010; Eide et al., 2017; Millero, 2013), thus it is selected for evaluating the product's regional applicability.
The spatial patterns of model-estimated isotope values (δ13Cest) along cruises 33MW19930704 and 33RO20050111 effectively captured the distributional characteristics of the observed values (δ13Cobs) (Fig. 4b vs. 4c). The reconstructed δ13CDIC product exhibited a very low transect-mean bias of −0.007 ‰ with a standard deviation of 0.082 ‰ (Fig. 4d), highlighting the product's reliability in estimating the spatial distribution of δ13CDIC. Spatially, the discrepancies are very small in most regions of the transect, but relatively larger in subpolar regions (around 50° S or 50° N) and vertically in the upper 500 m and below 3000 m (Fig. 4d).
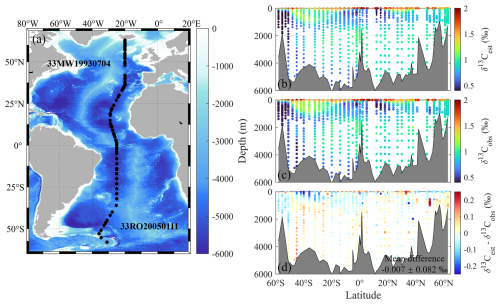
Figure 4(a) Station locations of the independent test cruises 33MW19930704 and 33RO20050111; Depth profile of (b) model-estimated (δ13Cest) and (c) in-situ observed (δ13Cobs) δ13CDIC along cruises 33MW19930704 and 33RO20050111, and (d) spatial distribution of mean bias error (MBE) between δ13Cest and δ13Cobs for the two cruises. Positive MBE values (red) denote product overestimation, while negative values (blue) indicate underestimation relative to observation. The overall mean difference is −0.007 ± 0.082 ‰.
3.3 Evaluation of the product's uncertainty
The uncertainty of the reconstructed δ13CDIC product was estimated by propagating uncertainties from three primary sources: measurement (uobs), input variables (uinputs) and mapping (umap). Detailed calculations are described in Sect. 2.5 of the Methods. To ensure a conservative estimate, we adopted a uniform uobs of 0.07 ‰ for all data points. umap is primarily quantified using the GPR model's posterior predictive standard deviation (i.e., the square root of the posterior predictive variance), which reflects the model's inherent uncertainty in mapping environmental predictors to δ13CDIC and captures both model variability and generalization-related uncertainty to unobserved regions. The mean value of this GPR-derived umap is 0.08 ‰. Following previous literature (Wu et al., 2025; Roobaert et al., 2024; Sharp et al., 2022), umap can alternatively be estimated as the RMSE between δ13Cest and δ13Cobs in the training set. We additionally report this RMSE-based metric as a reference for compatibility with existing studies, and notably, it yields a quantitatively similar value of 0.08 ‰ to the GPR-derived umap. uinputs was quantified via Monte Carlo simulation, considering contributions from seven environmental variables: T, S, AOU, N, Si, DIC, xCO2. Notably, uncertainties from input variables had a nearly negligible impact, with uinputs estimated at 0.009 ‰ . The uinputs contribution from individual input variables was decomposed as follows: temperature (4.96 × 10−5 ‰), salinity (3.62 × 10−4 ‰), nitrate (0.004 ‰), silicate (0.002 ‰), DIC (0.005 ‰), AOU (0.004 ‰), and xCO2 (6.52 × 10−4 ‰). Overall, the reconstructed δ13CDIC product exhibits an average uncertainty of 0.11 ‰ for the entire Atlantic Ocean. This uncertainty is considered reasonable given the conservative estimation approach, highlighting the product's reliability for δ13CDIC characterization.
3.4 Additional validation using data extracted from a numerical model environment
To address the inherent limitation of observational δ13CDIC data (sparse and unevenly distributed across space and time), we conducted supplementary numerical model-based validation to verify the GPR model's reliability. This validation utilized well-validated ocean carbon isotope simulation data from Claret et al. (2021), which includes global monthly outputs (1990–2002) with a 1° × 1° horizontal grid and 50 depth layers (Appendix A).
First, we focused on the Northeast Atlantic (30° W–5° W, 5° S–66° N) and downsampled the data to simulate observational sparsity. Limited by computational resources, we selected data from 1991 to 1995 (60 months) for validation, with 1993 (12 months) designated as an independent test set to assess generalization (Appendix A1.2). The GPR model adopted the same configuration as the observational-based reconstruction (Matern kernel, 10-fold cross-validation for hyperparameter tuning). Validation results demonstrated exceptional performance. For raw δ13CDIC values, R2 exceeded 0.99 and RMSE was less than 0.015 ‰ across the training set, independent test set, and additional prediction set (Figs. A1 and A2 in Appendix A1.3). Additionally, anomaly-based R2 (quantifying deviations from local climatological means) ranged from 0.67 to 0.90, confirming the model's ability to capture biogeochemically meaningful spatiotemporal variations independent of large-scale offsets. For detailed data processing, parameter settings, and full validation results, refer to Appendix A1.
Second, to further enhance the credibility of the GPR model in real-world application scenarios, we supplemented an additional observation-constrained sparse validation based on the same numerical model dataset (Appendix A2). This validation strictly mimics the spatial locations of sparse oceanographic observations by subsampling model data exclusively at the exact locations of the 37 Atlantic cruises and GLODAP locations, while preserving cruise-specific uneven sampling patterns (full experimental design detailed in Appendix A2.1). The results of this supplementary validation also showed robust performance. The training set achieved an R2 of 0.99 and an RMSE of 0.009 ‰, the independent test set yielded an R2 of 0.99 and an RMSE of 0.025 ‰, and the reconstructed values at GLODAP locations were highly consistent with the model's native values (R2=0.95, RMSE = 0.080 ‰) (Figs. A3 and A4, see Appendix A2.3 for details).
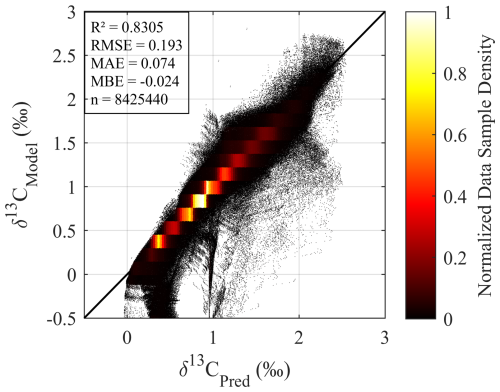
We further conducted a supplementary full Atlantic domain generalization test to verify the model's ability to reproduce large-scale δ13CDIC patterns across the entire basin. Using the model trained on observation-matched sparse data, we predicted gridded δ13CDIC values for the full Atlantic domain, with results showing strong overall alignment with the native model values (R2=0.83, RMSE = 0.193 ‰ across 8.4 million valid samples, Fig. A5 in Appendix A3). This test provides additional evidence that our approach can reliably leverage sparse observations to estimate basin-scale oceanographic δ13CDIC distributions.
The grid-based downsampling validation and the supplementary observation – constrained sparse validation complement each other, jointly verifying the GPR model's robustness in reconstructing δ13CDIC patterns from sparse data, whether under idealized sparse grid scenarios or real observational sparse scenarios. This dual validation framework provides a solid methodological foundation for subsequent observational-based reconstruction.
3.5 Reconstruction of δ13CDIC
Here, the trained GPR model is applied to GLODAPv2.2023 hydrographic data for δ13CDIC reconstruction in the Atlantic Ocean, spanning the 1980s to 2021 with a small number of 1972 data points. The GLODAPv2.2023 Atlantic dataset comprises 500 137 samples, of which 8941 contain acceptable δ13CDIC observations (quality flags = 2 or 6). A total of 124 643 δ13CDIC values can be reconstructed based on the availability of all required predictor variables (salinity, AOU, Nitrate, Silicate, and DIC). Among these, 68 435 reconstructions are considered high quality, as all six input variables have acceptable quality flags, which is approximately 7.65 times larger than the number of δ13CDIC observations. The remaining 56 208 reconstructions are based on input variables with unknown or lower quality and are thus assigned a quality flag of 3 (questionable). All 124 643 reconstructions, 13.9 times larger than the observation, are provided in the supplementary Dataset, of which the questionable δ13CDIC samples are provided for transparency but are not recommended for routine use without further quality assessment. The following presentations and analyses are restricted to the 68 435 acceptable samples to ensure the robustness of the results unless otherwise noted.
The observed and reconstructed datasets share an intersection of 5997 samples that have both acceptable δ13CDIC observations and acceptable input variables for reconstruction. These overlapping samples are used for direct model evaluation (Fig. 5a). The comparison between observed and reconstructed δ13CDIC values yields a high correlation coefficient (R2=0.89), with RMSE = 0.094 ‰, MAE = 0.067 ‰, and MBE 0.009 ‰. The mean difference between reconstruction and observation is −0.009 ± 0.097 ‰, confirming strong overall agreement. To further verify the statistical consistency between observed and reconstructed δ13CDIC values, we calculated Gaussian kernel density estimation (KDE) for both datasets (Fig. 5b). KDE is a non-parametric method for approximating probability density functions by summing Gaussian kernel functions centered at each data point (Silverman, 1986), producing a continuous estimate that is less sensitive to arbitrary bin edges. For this analysis, KDE was applied on a uniform grid spanning the full δ13CDIC range, with a bandwidth of 0.1 ‰ chosen to balance smoothing and resolution. No additional jittering was applied as the data contains sufficient variability to avoid overlapping artifacts. The KDE curves of observed (n = 8941, range from −0.37 ‰ to 2.37 ‰) and reconstructed (n = 68 435, range from −0.11 ‰ to 2.36 ‰) δ13CDIC are highly consistent, exhibiting nearly identical unimodal distributions, with a primary peak around 1 ‰. Both curves decline rapidly to near zero at the extremes (below ∼ 0 ‰ and above ∼ 2 ‰), reflecting minimal occurrence of δ13CDIC values in these ranges. This close alignment confirms the reconstructed δ13CDIC from GPR model accurately captures the central tendency, overall shape, and range of the observed δ13CDIC distribution.
The reconstructed dataset, which has a much larger sample size and spatially continuous coverage, exhibits minimally higher probability density than observations for δ13CDIC values between approximately 0.8 ‰ and 1.1 ‰, alongside a slightly sharper peak and reduced variance. This subtle distinction is a natural consequence of model smoothing intrinsic to GPR's Gaussian kernel framework and the mitigation of sampling noise in the dense, continuous reconstruction. By weighting neighbouring data points to produce spatially consistent predictions, GPR avoids overfitting to sparse extreme values often linked to transient perturbations or observational noise, an outcome aligned with the study's objective of generating a robust, representative δ13CDIC distribution. Overall, the KDE analysis, based on a systematic bandwidth selection and grid evaluation, underscores the strong statistical congruence between observed and reconstructed δ13CDIC, with the minor differences being negligible relative to the model's success in replicating the core characteristics of the observed distribution.
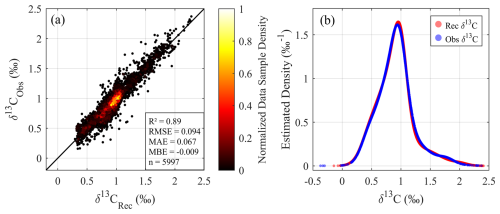
Figure 5Comparison of observed and reconstructed δ13CDIC values derived from the GLODAPv2.2023 dataset. (a) Density scatter plot of observed versus reconstructed δ13CDIC values, using only the intersection of data points with acceptable quality available in both observed and reconstructed datasets (n=5997). (b) Gaussian kernel density estimations (KDEs) for a comprehensive evaluation of observed and reconstructed δ13CDIC values. In panel (b), the blue curve represents all acceptable δ13CDIC observations (n=8941), while the red curve indicates all acceptable reconstructed δ13CDIC values (n=68 435), generated by GPR model trained on the GLODAPv2.2023 dataset. The KDEs reveal the overall distribution shape, highlight differences in peak height and spread, and provide a smooth, continuous representation of probability density.
The GLODAPv2.2023 Atlantic Ocean dataset includes approximately 20 583 stations in total (Fig. 6a). Among these, only about 732 stations have δ13CDIC observations (blue points in Fig. 6b), collected over the four decades, reflecting the limited spatial and temporal extent of direct observations due to the high cost and complexity of δ13C sampling and analysis. Using the proposed GPR model, δ13CDIC values have been reconstructed at roughly 4182 stations (red points in Fig. 6b). This reconstructed dataset significantly expands both temporal and spatial coverage of δ13CDIC in the Atlantic Ocean, enabling more detailed and extensive biogeochemical analyses across the Atlantic.
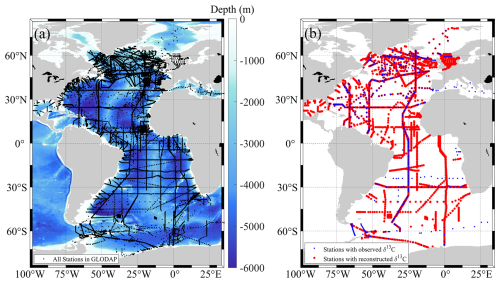
Figure 6Station maps for the GLODAPv2.2023 Atlantic Ocean dataset. (a) Locations of all stations included in the dataset. (b) Stations with observed (blue dots) and reconstructed (red dots) δ13CDIC.
To evaluate the spatial and temporal coverage expansion of the reconstructed δ13CDIC dataset, the data distributions are compared along longitude, latitude, year, and depth (Fig. 7). Along longitude (Fig. 7a), the number of reconstructed δ13CDIC increased several times compared to the observed, and also extended into west of 70° W and east of 0° E where there are almost no direct δ13CDIC observations. The latitudinal distribution (Fig. 7b) shows notable improvements in both hemispheres. The reconstructed dataset greatly enhanced the temporal coverage (Fig. 7c). There is little to no reconstructed data before 1980, between 1984 and 1987, and in 1995. The number of reconstructed data increases significantly after the late-1980s, reaching a peak in 2003, (Fig. 7c). Vertically, the data number of reconstructed δ13CDIC increased substantially throughout the water column, although numbers of both reconstructed and observed δ13CDIC data show exponential decrease from surface to deep water (Fig. 7d).
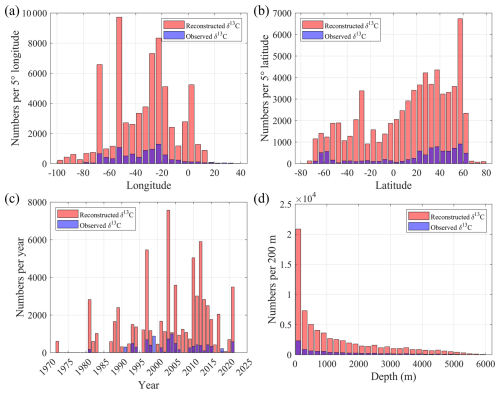
Figure 7Comparison of the spatial, temporal, and vertical distributions of observed and reconstructed δ13CDIC data in the Atlantic Ocean. The number of δ13CDIC samples is shown by (a) longitude (per 5° bin), (b) latitude (per 5° bin), (c) year (from 1972 to 2021), and (d) depth (per 200 m interval). Red bars represent reconstructed δ13CDIC, and blue bars indicate observed δ13CDIC.
Overall, the reconstructed δ13CDIC dataset consistently demonstrated good accuracy, high correlation coefficient, low RMSE, MAE, and minimal bias across diverse environmental conditions. Spatially, the reconstruction significantly expands coverage across longitude, latitude, and depth, particularly in undersampled regions such as the South Atlantic and deeper ocean. Temporally, it enhances data availability, filling many before 1990 and after 2015, despite some limitations tied to input variable availability.
Additionally, a spatially continuous 3D (1° × 1° horizontal resolution, 33 standard depth layers) climatological δ13CDIC dataset for the Atlantic Ocean was generated using the validated GPR framework. This product was built on gridded environmental predictors (temperature, salinity, oxygen (used to calculate AOU), nitrate, silicate, and DIC) from the GLODAPv2 1° × 1° global interior ocean mapped climatology (Lauvset et al., 2016) and monthly mean atmospheric xCO2 spanning 1972–2013. The resulting product aligned with the coordinate system and structural conventions of the GLODAPv2 gridded framework, ensuring full interoperability with mainstream ocean biogeochemistry research workflows. It is specifically designed to address the critical limitations of discrete site-level δ13CDIC observations for basin-scale spatial analyses. This gridded product is archived alongside our other datasets in the Zenodo repository (https://doi.org/10.5281/zenodo.18481145, Gao et al., 2025).
3.6 Potential Applications
The improved spatial, temporal, and vertical coverage of the reconstructed δ13CDIC dataset potentially contributes to the biogeochemical research and to a deeper understanding of carbon cycling processes in the Atlantic Ocean. Two specific applications are given as examples here: (1) examining the spatial patterns and decadal variability of surface δ13CDIC (≤ 10 m), and (2) evaluating its ability to resolve vertical profiles, thereby providing insights into subsurface carbon dynamics.
To facilitate a more comprehensive understanding of the spatial and temporal dynamics of surface δ13CDIC in the Atlantic, Fig. 8 illustrates how the reconstruction improves the spatial representativeness and temporal continuity of surface δ13CDIC. The expanded spatial coverage shown in Fig. 8a increases the representativeness of latitudinal sample bins, reducing the influence of sparsely sampled outliers on bin means (Fig. 8b). For instance, observational data in the 5–10° S band during the 2000s show an anomalously low mean that is likely driven by a few isolated observations. In contrast, the reconstructed dataset, by supplying a larger and spatially coherent sample set, yields a mean that is more consistent with adjacent latitude bands. This effect should be understood as a reduction of sampling-driven discontinuities and noise rather than as an artificial suppression of genuine signals. Both observed and reconstructed data indicate a basin-scale decline in surface δ13CDIC from the 1980s to the 2010s, most pronounced in tropical and subtropical regions (approximately 35° S–35° N), consistent with the expected Suess effect from increasing fossil-fuel CO2 (Keeling, 1979). In contrast, the mid- to high-latitude regions exhibit more complex variability, likely reflecting the influence of regional circulation, water mass formation, and subduction processes that modulate isotopic signals on multi-decadal timescales.
To facilitate a clear and intuitive assessment of decadal distributions of observed and reconstructed surface δ13CDIC, we employ KDE. KDE results presented in Fig. 8c are computed using all acceptable observed and reconstructed surface δ13CDIC samples within each decade and are intended to reveal the overall distributional characteristics and their evolution across decades and complement the latitude-binned means shown in Fig. 8b. In general, both observed and reconstructed KDE curves confirm the decadal downward shifts of δ13CDIC from the 1980s to the 2010s, reinforcing the notion of a basin-wide isotopic response to anthropogenic carbon inputs. And the observed and reconstructed KDE curves within each decade fall within comparable value ranges and exhibit similar peak positions, underscoring the consistency between the two datasets. Notable discrepancies arise in specific decades. For instance, in the 1980s the reconstructed KDE curve shows a dominant peak near 2 ‰ and a secondary peak near 1.7 ‰. The latter closely corresponds to the single, broader maximum in the observational KDE curve. This discrepancy is likely attributable to the sparse observational coverage during this decade (Fig. 8a), which limits the representativeness of the latitudinal distribution. In the 2000s, the reconstructed KDE curve exhibits slightly greater density at the lower end of the common range (∼ 0.5 ‰–2.0 ‰) than the observational KDE curve, producing a modest negative shift in central tendency while maintaining a comparable overall span. This shift is primarily attributable to a disproportionate increase in reconstructed data between 50 and 65° N, which modifies the density structure of the distribution. These examples highlight that the reconstruction not only reproduces the principal features of the observational distributions but also mitigates the limitations imposed by sparse or uneven sampling. The reconstructed KDE curves are generally smoother and more spatially coherent than the raw observational KDE curves. The smoother and more coherent appearance of the reconstructed KDE curves reflect the underlying basin-scale δ13CDIC structure. This enhanced spatial consistency allows the basin-wide decadal shift toward lower δ13CDIC values to emerge more clearly by reducing the influence of uneven sampling. As a result, the reconstruction provides a more representative depiction of δ13CDIC variability across latitude and time, improving the statistical robustness and interpretability of the inferred distributional changes.
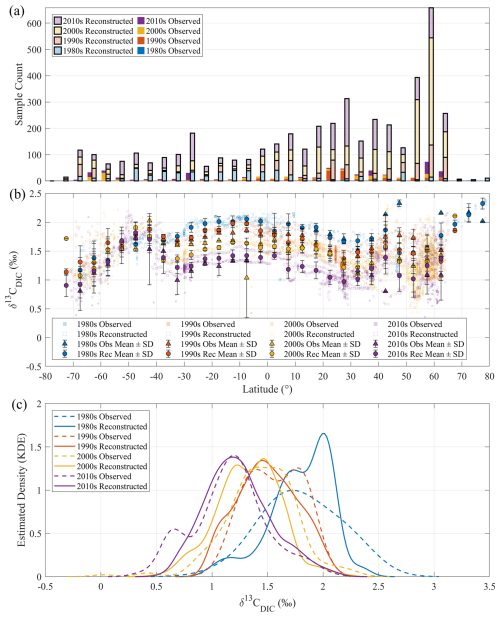
Figure 8Surface distribution of observed and reconstructed δ13CDIC in the Atlantic Ocean across four decades. (a) Number of observed and reconstructed δ13CDIC samples within each 5° latitude band for the 1980s, 1990s, 2000s, and 2010s. For each latitude band, stacked bars represent decadal sample counts. Solid (opaque) bars indicate observations, and semi-transparent bars with black outlines indicate corresponding reconstructions. (b) Surface δ13CDIC distributions (depth ≤ 10 m) along latitude for the 1980s, 1990s, 2000s, and 2010s. Colored scatter points indicate individual δ13CDIC values from observations (circles) and reconstructions (dots), while the black markers with corresponding face color and error bars represent the mean ± standard deviation of δ13CDIC within each 5° latitude bin, triangles for observed data and diamonds for reconstructed values. (c) Gaussian kernel density estimates (KDEs) of surface δ13CDIC for each decade (1980s–2010s), using dashed lines for observations and solid lines for reconstructions. These KDEs reveal the decadal evolution and overall distributional characteristics of δ13CDIC in both datasets.
Besides horizontal distributions, the reconstructed δ13CDIC dataset also provides valuable insights into vertical variability. The depth profiles along the North Atlantic A16N transect in 1993, 2003, 2013, and 2023 (Fig. 9) show that the reconstruction substantially improves vertical resolution and continuity, especially for years with sparse measurements. For instance, the δ13CDIC samples were increased from 493 to 1618 in 1993, 38 to 2395 in 2003, and 473 to 2787 in 2013, respectively, enhancing data coverage across depths and latitudes, facilitating the detection of temporal trends associated with ocean carbon uptake and redistribution (Fig. 9). The reconstructed δ13CDIC values for 1993, 2003, and 2013 were generated by applying the pre-trained model to input variables from the GLODAPv2.2023 Atlantic dataset. Notably, as the observational data along A16N in 2023 not included in the GLODAPv2.2023 dataset, we used the same pre-trained and validated model, relying solely on the 2023 observational input variables to produce the reconstructed values. The close alignment between 2023's reconstructed and observed data (Fig. 9d vs. 9h) not only reflects the model's reliability but also validates its ability to generalize, strengthening confidence in reconstructions for years with sparse measurements (e.g., 2003 with only 38 observations).
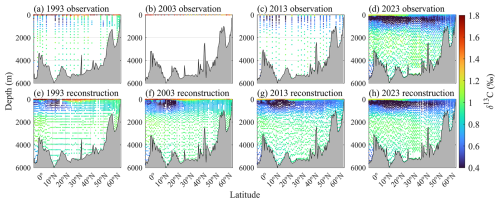
Figure 9Enhancement of spatial resolution for δ13CDIC through GPR model reconstruction along North Atlantic A16N transect. Top panels (a)–(d) present in-situ observed δ13CDIC from cruises in (a) 1993 (n=493), (b) 2003 (surface-only, n=38), (c) 2013 (n=473), and (d) 2023 (n=3460). Bottom panels (e)–(h) show corresponding reconstructed δ13CDIC with systematic filling of spatial gaps: (e) the number of reconstructed δ13CDIC for cruise A16N in 1993 is 1618; (f) full-depth reconstruction for cruise A16N in 2003 (n=2395); (g) 2013 reconstruction (n=2787), highlighting improved coverage compared to limited observations (n=473); (h) the reconstruction of cruise A16N in 2023 (n=3030) validated against high-density observations. Reconstruction effectively resolves historical sampling sparsity, yielding consistent spatial resolution comparable to contemporary DIC measurements.
The enhanced vertical profiles provided by the reconstructed δ13CDIC dataset also enable more refined investigations into the physical and biological controls governing carbon isotope variability in the ocean interior. Joint interpretation of δ13CDIC with nutrients such as phosphate allows for the identification of water mass structures and biological removal and addition, shedding light on processes such as organic carbon export, remineralization, and the coupling between biological productivity and ocean circulation (e.g., Gruber et al., 1999; Eide et al., 2017). These capabilities improve the isolation of the vertical imprint of the Suess effect and facilitate the reconstruction of preindustrial δ13CDIC, both of which are critical for assessing anthropogenic perturbations to the marine carbon cycle (Olsen and Ninnemann, 2010).
δ13CDIC also can be used to estimate biological carbon export and net community production (NCP) (Quay et al., 2009; Yang et al., 2019). Accurate NCP estimation from δ13CDIC requires knowledge of both the physical supply of DIC and δ13CDIC, typically represented by the subsurface to surface δ13CDIC gradient. Sparse historical δ13CDIC data can bias this estimate, whereas the reconstructed dataset provides a continuous field that reduces gaps and noise, enabling more reliable NCP calculations.
Importantly, the observation-constrained reconstructed δ13CDIC fields fill longstanding gaps in global datasets, providing a robust basis for the validation of Earth system models (e.g., Schmittner et al., 2013; Sonnerup and Quay, 2012; Claret et al., 2021). The improved coverage helps reconcile discrepancies between modeled and observed δ13CDIC distributions, particularly in data-sparse regions such as the deep ocean and the Southern Hemisphere. In particular, at the boundary of two water masses, the high resolution δ13CDIC distribution helps to validate high-resolution global physical and biogeochemical model predictions and more effectively study carbon cycle at such boundary zones.
Overall, the reconstruction effectively addresses key limitations of the observational δ13CDIC record, particularly data sparsity and sampling bias, providing a more continuous and spatially balanced dataset. This enables clearer identification of large-scale latitudinal gradients, decadal trends, and regional anomalies, offering a robust foundation for interpreting long-term carbon cycle dynamics.
3.7 Challenges and Limitations
Despite enhancing spatial resolution, reconstructing grid δ13CDIC with low uncertainty remains challenging compared to other carbonate system variables (e.g., DIC, pCO2, TA), primarily due to limited historical observation coverage. While the dataset achieves an overall mean bias of −0.007 ‰, notable regional discrepancies persist, particularly in subpolar regions, where biases of some samples exceed 0.15 ‰. These anomalies may be attributed to highly nonlinear interactions between δ13CDIC and biogeochemical processes, such as upwelling-induced isotope fractionation and biological carbon pump effects (Gruber et al., 1999). Additionally, although input variables (T, S, AOU, N, Si, DIC, xCO2) were validated to have a small impact on reconstruction uncertainty (0.009 ‰), the exclusion of potential covariates, such as chlorophyll a, wind speed, or nutrient gradients, may limit constraint precision.
The reconstruction approach is also subject to inherent limitations, such as spatial interpolation assumptions, uncertainty propagation, and temporal variability constraints. The GPR model assumes stationarity of δ13CDIC spatial covariance, a simplification that may fail in regions with abrupt bathymetric changes (e.g., the Mid-Atlantic Ridge). Such nonstationarity may introduce systematic errors in reconstructed δ13CDIC gradients, particularly across topographic features that influence ocean circulation and carbon transport. Cumulative uncertainties from measurement (0.07 ‰), mapping (0.08 ‰), and input variables (0.009 ‰) yield an overall uncertainty of 0.11 ‰, which may obscure small-scale δ13CDIC signals. This limitation restricts the dataset's utility for resolving fine-scale temporal trends or localized isotopic anomalies. Furthermore, the temporal sparsity of historical predictor observations restricts the resolution of interannual δ13CDIC variability. The application of reconstructed data to estimate the Suess effect using extended Multilinear Regression (eMLR) methods in repeated hydrographic transects requires careful, case-specific consideration. Our preliminary analysis confirms that sample density significantly affects the anthropogenic δ13CDIC changes estimated via the eMLR method. Results derived from the sparse downsampling δ13CDIC data deviate from those obtained using full observational datasets, while the anthropogenic δ13CDIC changes calculated from our reconstructed high-density δ13CDIC data show strong consistency with the full-observation benchmark. However, the magnitude of decadal-scale Suess effect (∼ 0.2 ‰) is comparable to the overall uncertainty of the reconstructed δ13CDIC values (0.11 ‰). Thus, further rigorous and more comprehensive assessment of these uncertainties is critical to ensure the reliability of decadal-scale isotopic trend estimates derived from eMLR analyses, and to avoid biased or erroneous conclusions.
To address these challenges and limitations, future work will be focused on enhancing mechanistic insights into how δ13CDIC relates to environmental factors. Subregional partitioning strategies and variable selection methods may be developed to improve model accuracy, while process-based knowledge also should be integrated to identify optimal predictor variables and refine regionalization schemes for biogeochemical heterogeneity. Additionally, uncertainty mitigation techniques will be coupled with emerging high-resolution observations from autonomous platforms, which will be leveraged to refine spatial resolution to fixed grids. Furthermore, physics-informed reconstruction frameworks (e.g., Bennington et al., 2022) have the potential to extend δ13CDIC records across pre- and post-satellite eras using proxy variables for historical biological productivity, improving temporal consistency and Suess effect estimation via eMLR methods by reducing uncertainties in decadal-scale trend analyses.
All products described in this work are publicly archived in the Zenodo data repository under DOI: https://doi.org/10.5281/zenodo.18481145 (Gao et al., 2025). This archive contains three complementary, community-standard compliant datasets: (1) a GPR-reconstructed δ13CDIC dataset based on the GLODAPv2.2023 Atlantic Ocean subset, with full interoperability with standard GLODAPv2 Atlantic Ocean subset workflows; (2) a comprehensive quality-controlled compilation of δ13CDIC observational data from 51 Atlantic cruises, supporting method reproducibility and independent validation; and (3) the 1° × 1° climatological gridded 3D δ13CDIC dataset aligned with the GLODAPv2.2016b mapped climatology (Lauvset et al., 2016), for spatially continuous basin-scale analyses.
The MATLAB code used to process the data and create the figures included in this paper is provided at https://doi.org/10.5281/zenodo.19244239 (huigao109, 2026).
This study reconstructs Atlantic Ocean δ13CDIC using a GPR model trained on 37 secondary quality-controlled cruises selected from 51 compiled historical datasets, including a high-resolution 2023 A16N transect. GPR was chosen for its ability to capture nonlinear relationships, outperforming other machine-learning models in preliminary tests (lowest RMSE and highest R2). Additionally, the method's robustness was validated using numerical model data (Claret et al., 2021), with both grid-based downsampling and observation-constrained sparse sampling validation frameworks, confirming not only the GPR model's stability but also the high reliability in δ13CDIC reconstruction. The trained GPR model achieved an average bias of only −0.007 ± 0.082 ‰ and an overall uncertainty of 0.11 ‰, with error contributions from measurement, mapping and input-variable.
Applying GLODAPv2.2023 Atlantic dataset as predictors, the reconstruction expanded the acceptable δ13CDIC samples from 8941 to 68 435, representing a substantial 7.65-fold increase compared to the original dataset. This extensive dataset significantly improves spatial coverage in longitude, latitude, and depth and enhances temporal continuity of δ13CDIC observations over the past four decades.
Multiple pieces of evidence attest to reliability and superiority of the reconstructed dataset. Statistical diagnostics demonstrate strong model skill in both cross-validation and independent testing stages, as well as reconstruction stage. The reconstruction preserves decadal variability at the sea surface δ13CDIC and in depth-profiles and agrees with high-density contemporary observations such as the 2023 A16N transect. Distributional metrics, exemplified by smoothed and stable KDE curves, suggest that the reconstruction mitigates sampling noise while retaining meaningful spatiotemporal signals.
The broad spatial coherence and high coverage of the reconstructed δ13CDIC enable systematic analysis of large-scale gradients, detection of regional anomalies, and investigation of tracer relationships with nutrients such as phosphate, supporting a wide range of carbon cycle studies. It also provides a valuable baseline for evaluation of Earth system models, for improving estimates of preindustrial and anthropogenic δ13CDIC, and for extending isotopic records used in climate reanalysis.
Despite these advances, some challenges and limitations remain. Regional biases persist, likely reflecting highly nonlinear biogeochemical interactions and the GPR assumption of spatial covariance stationarity, which may break down over complex bathymetry. Small-scale features can be obscured by cumulative uncertainty propagation and temporal inconsistencies persist in certain derived quantities. Future efforts will be needed on assimilating higher-resolution observations to enhancing the mechanistic understanding of relationships between δ13CDIC and environment factors, developing tailored subregional modelling frameworks and exploiting advanced machine-learning techniques to capture nonstationary spatial features. These efforts will refine spatial and temporal fidelity, reduce uncertainties and gain deeper mechanistic insight into ocean carbon cycle dynamics.
The main text focuses on reconstructing δ13CDIC in the Atlantic Ocean using sparse observational data. However, validating machine learning methods solely with observations faces inherent limitations: observational data are spatially-temporally discrete, unevenly distributed, and lack an explicit reference distribution for rigorous accuracy assessment. To address this gap, we conducted supplementary numerical model-based validation using well-validated ocean carbon isotope simulation data (Claret et al., 2021). This validation aims to evaluate the GPR model's ability to reconstruct 4D (longitude-latitude-depth-time) δ13CDIC patterns from sparse, perturbed data and complement observational validation to strengthen the methodological rigor.
The original simulation dataset from Claret et al. (2021) features monthly outputs from 1990 to 2002, a 1° × 1° horizontal grid (global longitude-latitude) and 50 depth layers covering the full water column. Notably, the dataset does not provide nitrate (N) or silicate (Si) data but includes phosphate (PO4) data. We conducted sensitivity tests on observational dataset to assess the impact of input variable selection, comparing two variable sets: one including longitude, latitude, depth, T, S, AOU, nitrate, silicate, DIC, and xCO2, and the other replacing N and Si with PO4. Results showed no significant performance differences between the two sets (the change of RMSE < 2 %), confirming that PO4 can effectively serve as a proxy for the missing nutrients.
A1 Grid-based Subsampling Validation (Idealized Sparse Scenario)
A1.1 Experimental Design and Dataset Partitioning
Due to the massive size of the original dataset (exceeding computational feasibility for standard Machine Learning workflows), we processed the data as follows: (1) We focused on the Northeast Atlantic (30–5° W, 5° S–66° N), a region with representative δ13CDIC spatial gradients and relevance to the main text's Atlantic-focused reconstruction. (2) To simulate observational sparsity, we subsampled the 1° × 1° grid at 5-grid intervals, resulting in a horizontal resolution of 5° × 5° (5 longitude points × 19 latitude points within the study region). (3) For the vertical dimension, we retained depth layers within the upper 1500 m, and subsampled 1 point every 4 layers, yielding 10 vertical layers. Temporally, we selected data from 1991 to 1995 (60 months) for validation, with 1993 (12 months) designated as an independent test set to assess generalization.
Each month's theoretical sample size after downsampling is 950, and after excluding grid cells with invalid data, the monthly effective sample size is 799. Over 60 months, the total sample size is 47 940. Given this scale, we randomly selected of the data (15 980 samples) for model training, validation, and independent testing, with the remaining (31 960 samples) as an additional prediction set to further verify generalization to unseen 4D grid points.
A1.2 Model Configuration and Evaluation Metrics
The GPR model used in this validation adopted the same configuration as the main text: Matern kernel, and hyperparameter tuning via 10-fold cross-validation within the training set. This consistency ensures the validation results are directly comparable to the main text's observational-based performance. For dataset partitioning, the training set comprised 80 % of the subsampled data covering 1991–1992 and 1994–1995 (excluding 1993), while the internal validation set included 20 % of the subsampled data from the same non-1993 years as the training set and was used for hyperparameter tuning. The independent test set included all 1993 data within the subsampled pool to ensure temporal independence from training/validation data, and the additional prediction set consisted of the remaining of the full downsampled data (31 960 samples) spanning 1991–1995 to verify reconstruction of dense 4D grids. Consistent with the main text, we used RMSE and R2 to assess point prediction accuracy. We further introduced anomaly-based R2 (), which is calculated from anomalies relative to the local mean or climatology, where the local climatological mean denotes the 5-year monthly average of δ13CDIC at each longitude-latitude-depth grid, to quantify the model's ability to capture biogeochemically meaningful deviations from large-scale background trends. By isolating these deviations from the climatological baseline, this approach reduces the influence of large-scale offsets and thereby provides a more realistic assessment of the model's capacity to reproduce spatial–temporal variations.
A1.3 Validation Results
The GPR model achieved exceptional performance in the model-based validation, confirming its robustness for 4D δ13CDIC reconstruction. For raw δ13CDIC values (total signal), the internal validation set yielded R2>0.99 and RMSE = 0.006 ‰, while the independent test set showed R2>0.99 and RMSE = 0.012 ‰ (Fig. A1), and the additional prediction set exhibited R2>0.99 and RMSE = 0.013 ‰ (Fig. A2). Critically, the anomaly-based R2 values further demonstrated the model's capacity to capture ecologically relevant variability: the training set reached an of 0.90, the temporally independent test set achieved an of 0.67, and the additional prediction set (unseen 4D grid points) showed an of 0.84. These results indicate the model nearly perfectly reproduces the absolute δ13CDIC values from the Claret model, while also effectively reconstructing biogeochemically meaningful anomalies such as seasonal cycling and regional biogeochemical deviations independent of large-scale climatological offsets. The consistent anomaly-based performance across the temporally unseen independent test set and the spatiotemporally dense, unseen additional prediction set confirms robust generalization to unobserved 4D δ13CDIC patterns. The slightly lower of 0.67 for the independent test set may stem from the limited sample size. Using only of the data rather than the full dataset likely insufficiently captures the unique spatiotemporal variability of 1993. Notably, the higher of the additional prediction set implies that a larger sample size may better reflect the spatiotemporal characteristics of δ13CDIC. However, in this paper, our focus is on large scale features over long timescales. We will examine this issue in future studies.
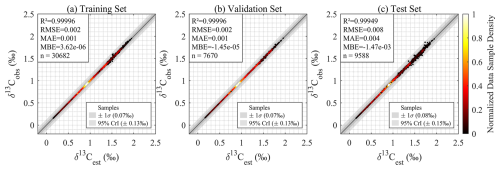
Figure A1GPR model evaluation for numerical model δ13CDIC reconstruction: Density scatter plots comparing GPR model-estimated (δ13Cest) versus numerical model outputs δ13CDIC (δ13Cobs) values during (a) training, (b) validation, and (c) independent testing.
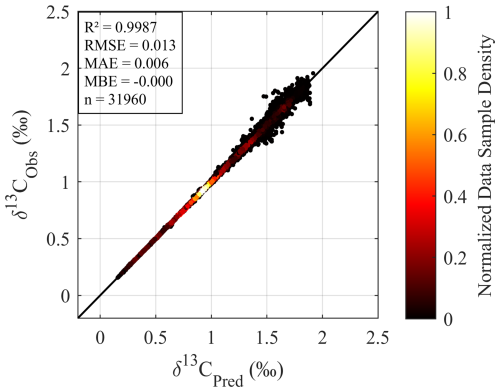
Figure A2Density scatter plot comparing numerical model outputs δ13CDIC (δ13CObs) versus predicted δ13CDIC (δ13CPred) values.
This model-based validation complements the observational validation by addressing two critical limitations of sparse, uneven coverage data. The exceptional performance (R2>0.99, RMSE < 0.015 ‰) confirms that the GPR model can accurately reconstruct 4D δ13CDIC patterns, validating its performance. In the meantime, meaningful anomaly-based R2 values ranging from 0.67 to 0.90 validate the model's capacity to capture ecologically and biogeochemically relevant variations beyond simply reproducing large-scale background trends. This validation strengthens confidence that the reconstructed dataset is not constrained by methodological flaws. Instead, its limitations stem from inherent observational uncertainties such as sparse sampling and unmodeled biogeochemical variability, ensuring the dataset reliably captures the spatial-temporal dynamics of δ13CDIC in the Atlantic Ocean.
A2 Observation-constrained Sparse Sampling Validation (Real-world Observational Scenario)
A2.1 Experimental Design Aligned with In-situ Observational Framework
To further enhance the credibility of the GPR model and the reconstructed dataset, and to address the concern that idealized grid-based subsampling may not fully reflect real-world observational scenarios, we supplemented an additional model-based validation that strictly mimics the spatial-temporal characteristics of sparse oceanographic observations. This validation focuses on evaluating the model's ability to learn from observation-like sparse data and generalize to target locations (e.g., GLODAP sampling locations), which is directly aligned with the core application scenario of the main text's δ13CDIC reconstruction work.
The validation design strictly follows the real observational framework: (1) We subsampled the Claret et al. (2021) model data exclusively at the exact spatial locations of the 37 Atlantic cruises used in the main text's training, validation, and test sets, as well as the GLODAP sampling locations employed for prediction. Given that the model data only spans 1990–2002, we mapped observational data from other decades (e.g., 2000–2009 observations to 1990–1999 model data) to maintain a sample size comparable to the full observational dataset. This design exactly replicates the sparse, heterogeneous distribution of real oceanographic observations, eliminating the idealized grid-based subsampling bias. (2) The GPR model adopted the same parameter settings as the aforementioned grid-based validation, ensuring consistency and comparability of results across different validation scenarios.
A2.2 Model Configuration and Evaluation Metrics
Consistent with the previous validation, we used R2, RMSE, MAE and MBE to assess the accuracy of point predictions for the training, validation, and independent test sets. For the GLODAP location prediction phase, we additionally performed Gaussian kernel density estimation (KDE) to evaluate the statistical congruence between reconstructed and native model δ13CDIC distributions.
A2.3 Validation Results
The results of this observation-constrained sparse validation demonstrate robust performance of the GPR model (Fig. A3). The training set achieves an R2 of 0.999 and an RMSE of 0.009 ‰, the validation set shows an R2 of 0.9985 and an RMSE of 0.012 ‰ , and the independent test set yields an R2 of 0.9950 and an RMSE of 0.025 ‰. These metrics confirm that the model can learn robustly from sparse, observation-like data and generalize effectively to unseen cruises, which is consistent with the performance of the grid-based validation and further verifies the model's stability.
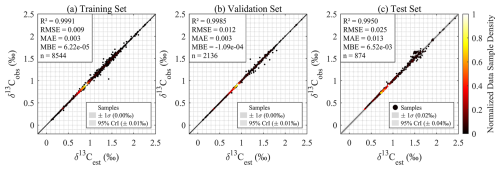
Figure A3GPR model evaluation for numerical model δ13CDIC reconstruction (observation-constrained sparse sampling): Density scatter plots comparing GPR model-estimated (δ13Cest) versus numerical model outputs δ13CDIC (δ13Cobs) values during (a) training, (b) validation, and (c) independent testing.
For the prediction phase at GLODAP locations (Fig. A4), the reconstructed δ13CDIC values align closely with the model's native values, with an R2 of 0.95 and an RMSE of 0.080 ‰. High-density data points cluster tightly along the 1:1 line (Fig. A4a), indicating excellent consistency between the reconstructed results and the true model values. Some minor deviations from the 1:1 line are likely attributed to inherent uncertainties and biases in the numerical model itself, rather than the GPR model's performance. The KDE analysis (Fig. A4b) further underscores the strong statistical congruence between the numerical model δ13C values and the reconstructed estimates, with minor differences being negligible relative to the model's success in replicating the core characteristics of the native distribution.
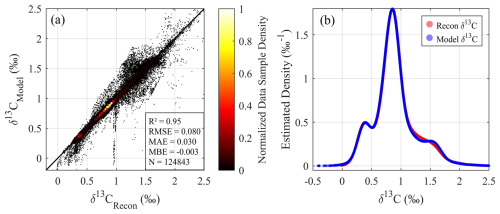
Figure A4Comparison of model and reconstructed δ13C values at GLODAP locations (observation-constrained sparse sampling): (a) Density scatter plot of model versus reconstructed δ13C values (n=124 843). (b) Gaussian kernel density estimations (KDEs) for a comprehensive evaluation of model and reconstructed δ13CDIC values.
A3 Supplementary Full Atlantic Domain Generalization Test
To further verify the GPR model's ability to reproduce the full spatial-temporal distribution of δ13CDIC across the entire Atlantic Ocean, we conducted a supplementary full-domain generalization test using the Claret et al. (2021) model data. Given the high computational memory demand of the original 1° × 1° horizontal resolution, 50 vertical levels full-domain dataset, we applied a sparse sampling strategy to maintain computational feasibility: we subsampled every other grid point in both the horizontal and vertical dimensions, and focused on the 1990–1999 period (120 months) for this test.
We used the identical GPR model trained on the observation-matched model data (Sect. A2), and predicted gridded δ13CDIC values across the entire Atlantic domain, then compared these predictions to the model's native “true” δ13CDIC values. The results show strong overall alignment between the reconstructed and native model values (Fig. A5), with an R2 of 0.8305, an RMSE of 0.193 ‰, an MAE of 0.074 ‰, and an MBE of −0.024 ‰ across 8 425 440 valid samples. High-density data points cluster tightly along the 1:1 line, confirming that the model can reliably reproduce the large-scale spatial-temporal patterns of δ13CDIC across the full Atlantic domain, and further demonstrating the model's strong generalization ability beyond the validation subsets.
This test provides additional evidence that our approach can effectively leverage sparse in-situ observations to estimate large-scale oceanographic δ13CDIC distributions. Limited by current computational capability, this test is a preliminary assessment of full-domain generalization performance; more comprehensive optimization and in-depth analysis of full-basin reconstruction will be conducted in future work.
A4 Conclusions of the Dual Validation Framework
This supplementary observation-constrained validation complements the previous grid-based validation, jointly reinforcing the reliability of the GPR model and the reconstructed data from two dimensions: “idealized grid sparse scenario” and “real observational sparse scenario”. The consistent excellent performance across both validation frameworks confirms that the GPR model can not only accurately reconstruct δ13CDIC patterns from sparse grid data but also reliably learn from real-world sparse observational data and generalize to target application locations. This dual validation provides solid methodological support for the rigor of the main text's δ13CDIC reconstruction work, further confirming that the reconstructed dataset can effectively address the limitations of sparse observational data and meet the needs of Atlantic carbon cycle research.
HG: Conceptualization, data curation, formal analysis, methodology, software, visualization, writing – original draft preparation, writing – review and editing. ZW: Validation, methodology, writing – review and editing. ZS: Validation, writing – review and editing. DC: Validation, methodology, writing – review and editing. MJ: Conceptualization, validation, writing – review and editing. WJC: Conceptualization, funding acquisition, writing – review and editing, project administration, supervision.
The contact author has declared that none of the authors has any competing interests.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. The authors bear the ultimate responsibility for providing appropriate place names. Views expressed in the text are those of the authors and do not necessarily reflect the views of the publisher.
The authors gratefully acknowledge all researchers, principal investigators, captains, and crew members who contributed their time at sea and in the laboratory to collect, analyze, and curate the datasets used in this study. We thank the providers and maintainers of the GLODAP, OCADS, and CCHDO databases, as well as the creators of the internally consistent δ13CDIC dataset for the North Atlantic Ocean (NAC13v1; Becker et al., 2016), for making their data openly available. The authors used ChatGPT for grammar and language polishing; all scientific content, interpretations, and conclusions are the sole responsibility of the authors.
This research is supported by the US National Science Foundation awards (OCE-2123768 and OCE-25A00158) to W.-J.C.
This paper was edited by Xingchen (Tony) Wang and reviewed by Patrick Rafter, Bin Lu, and one anonymous referee.
Alling, V., Porcelli, D., Mörth, C.-M., Anderson, L. G., Sanchez-Garcia, L., Gustafsson, Ö., Andersson, P. S., and Humborg, C.: Degradation of terrestrial organic carbon, primary production and out-gassing of CO2 in the Laptev and East Siberian Seas as inferred from δ13C values of DIC, Geochim. Cosmochim. Ac., 95, 143–159, https://doi.org/10.1016/j.gca.2012.07.028, 2012.
Becker, M., Andersen, N., Erlenkeuser, H., Humphreys, M. P., Tanhua, T., and Körtzinger, A.: An internally consistent dataset of δ13C-DIC in the North Atlantic Ocean – NAC13v1, Earth Syst. Sci. Data, 8, 559–570, https://doi.org/10.5194/essd-8-559-2016, 2016.
Bennington, V., Galjanic, T., and McKinley, G. A.: Explicit Physical Knowledge in Machine Learning for Ocean Carbon Flux Reconstruction: The pCO2 – Residual Method, J. Adv. Model Earth Sy., 14, e2021MS002960, https://doi.org/10.1029/2021MS002960, 2022.
Carter, B. R., Sharp, J. D., Dickson, A. G., Álvarez, M., Fong, M. B., García-Ibáñez, M. I., Woosley, R. J., Takeshita, Y., Barbero, L., Byrne, R. H., Cai, W., Chierici, M., Clegg, S. L., Easley, R. A., Fassbender, A. J., Fleger, K. L., Li, X., Martín-Mayor, M., Schockman, K. M., and Wang, Z. A.: Uncertainty sources for measurable ocean carbonate chemistry variables, Limnol. Oceanogr., 69, 1–21, https://doi.org/10.1002/lno.12477, 2024.
Claret, M., Sonnerup, R. E., and Quay, P. D.: A Next Generation Ocean Carbon Isotope Model for Climate Studies I: Steady State Controls on Ocean 13C, Global Biogeochem. Cy., 35, e2020GB006757, https://doi.org/10.1029/2020GB006757, 2021.
Deng, X., Li, Q., Su, J., Liu, C.-Y., Atekwana, E., and Cai, W.-J.: Performance evaluations and applications of a δ13C-DIC analyzer in seawater and estuarine waters, Sci. Total Environ., 833, 155013, https://doi.org/10.1016/j.scitotenv.2022.155013, 2022.
Eide, M., Olsen, A., Ninnemann, U. S., and Johannessen, T.: A global ocean climatology of preindustrial and modern ocean δ13C, Global Biogeochem. Cy., 31, 515–534, https://doi.org/10.1002/2016GB005473, 2017.
Esposito, M., Achterberg, E. P., Bach, L. T., Connelly, D. P., Riebesell, U., and Taucher, J.: Application of Stable Carbon Isotopes in a Subtropical North Atlantic MesocosmStudy: A New Approach to Assess CO2 Effects on the Marine Carbon Cycle, Front. Mar. Sci., 6, 616, https://doi.org/10.3389/fmars.2019.00616, 2019.
Gao, H., Cai, W., Jin, M., Dong, C., and Timmerman, A. H. V.: Ocean Ventilation Controls the Contrasting Anthropogenic CO2 Uptake Rates Between the Western and Eastern South Atlantic Ocean Basins, Global Biogeochem. Cy., 36, e2021GB007265, https://doi.org/10.1029/2021GB007265, 2022.
Gao, H., Jin, M., Zhao, H., Hussain, N., and Cai, W.: Using DIC-δ13C Pair to Constrain Anthropogenic Carbon Increase in the Southeastern Atlantic Ocean Over the Most Recent Decade (2010–2020), J. Geophys. Res.-Oceans, 129, e2024JC021586, https://doi.org/10.1029/2024JC021586, 2024.
Gao, H., Wu, Z., Sun, Z., Cai, D., Jin, M., and Cai, W.-J.: Reconstruction of δ13CDIC in the Atlantic Ocean, Zenodo [data set], https://doi.org/10.5281/zenodo.16907402, 2025.
Gregor, L. and Gruber, N.: OceanSODA-ETHZ: a global gridded data set of the surface ocean carbonate system for seasonal to decadal studies of ocean acidification, Earth Syst. Sci. Data, 13, 777–808, https://doi.org/10.5194/essd-13-777-2021, 2021.
Gruber, N., Keeling, C. D., Bacastow, R. B., Guenther, P. R., Lueker, T. J., Wahlen, M., Meijer, H. A. J., Mook, W. G., and Stocker, T. F.: Spatiotemporal patterns of carbon-13 in the global surface oceans and the oceanic suess effect, Global Biogeochem. Cy., 13, 307–335, https://doi.org/10.1029/1999GB900019, 1999.
Hughes, I. and Hase, T. P. A.: Measurements and their uncertainties: a practical guide to modern error analysis, New York, Oxford University Press, Oxford, 136 pp., ISBN 9780199566334, E-book ISBN 9780191576560, 2010.
huigao109: huigao109/ReC13_ML: ReC13_ML (v1.0), Zenodo [code], https://doi.org/10.5281/zenodo.19244239, 2026.
Keeling, C. D.: The Suess effect: 13Carbon-14Carbon interrelations, Environ. Int., 2, 229–300, https://doi.org/10.1016/0160-4120(79)90005-9, 1979.
Körtzinger, A., Quay, P. D., and Sonnerup, R. E.: Relationship between anthropogenic CO2 and the 13C Suess effect in the North Atlantic Ocean, Global Biogeochem. Cy., 17, https://doi.org/10.1029/2001GB001427, 2003.
Kroopnick, P.: The distribution of 13C in the Atlantic Ocean, Earth Planet. Sc. Lett., 49, 469–484, https://doi.org/10.1016/0012-821X(80)90088-6, 1980a.
Kroopnick, P.: Isotopic fractionations during oxygen consumption and carbonate dissolution within the North Atlantic Deep Water, Earth Planet. Sc. Lett., 49, 485–498, https://doi.org/10.1016/0012-821X(80)90089-8, 1980b.
Kroopnick, P.: The distribution of 13C of ΣCO2 in the world oceans, Deep-Sea Res. Pt. A, 32, 57–84, https://doi.org/10.1016/0198-0149(85)90017-2, 1985.
Kwon, E. Y., DeVries, T., Galbraith, E. D., Hwang, J., Kim, G., and Timmermann, A.: Stable Carbon Isotopes Suggest Large Terrestrial Carbon Inputs to the Global Ocean, Global Biogeochem. Cy., 35, e2020GB006684, https://doi.org/10.1029/2020GB006684, 2021.
Landschützer, P., Gruber, N. and Bakker, D. C. E. Decadal variations and trends of the global ocean carbon sink. Global Biogeochem. Cy. 30, 1396–1417, https://doi.org/10.1002/2015gb005359, 2016.
Lauvset, S. K. and Tanhua, T.: A toolbox for secondary quality control on ocean chemistry and hydrographic data, Limnol. Oceanogr. Meth., 13, 601–608, https://doi.org/10.1002/lom3.10050, 2015.
Lauvset, S. K., Key, R. M., Olsen, A., van Heuven, S., Velo, A., Lin, X., Schirnick, C., Kozyr, A., Tanhua, T., Hoppema, M., Jutterström, S., Steinfeldt, R., Jeansson, E., Ishii, M., Perez, F. F., Suzuki, T., and Watelet, S.: A new global interior ocean mapped climatology: the 1° × 1° GLODAP version 2, Earth Syst. Sci. Data, 8, 325–340, https://doi.org/10.5194/essd-8-325-2016, 2016.
Lauvset, S. K., Lange, N., Tanhua, T., Bittig, H. C., Olsen, A., Kozyr, A., Álvarez, M., Azetsu-Scott, K., Brown, P. J., Carter, B. R., Cotrim da Cunha, L., Hoppema, M., Humphreys, M. P., Ishii, M., Jeansson, E., Murata, A., Müller, J. D., Pérez, F. F., Schirnick, C., Steinfeldt, R., Suzuki, T., Ulfsbo, A., Velo, A., Woosley, R. J., and Key, R. M.: The annual update GLODAPv2.2023: the global interior ocean biogeochemical data product, Earth Syst. Sci. Data, 16, 2047–2072, https://doi.org/10.5194/essd-16-2047-2024, 2024.
Lima, I. D., Wang, Z. A., Cameron, L. P., Grabowski, J. H., and Rheuban, J. E.: Predicting Carbonate Chemistry on the Northwest Atlantic Shelf Using Neural Networks, J. Geophys. Res.-Biogeo., 128, e2023JG007536, https://doi.org/10.1029/2023JG007536, 2023.
Liu, B., Six, K. D., and Ilyina, T.: Incorporating the stable carbon isotope 13C in the ocean biogeochemical component of the Max Planck Institute Earth System Model, Biogeosciences, 18, 4389–4429, https://doi.org/10.5194/bg-18-4389-2021, 2021.
Millero, F. J.: Chemical Oceanography, 4th Edn., CRC Press, ISBN 9781466512498, https://doi.org/10.1201/b14753, 2013.
Olsen, A. and Ninnemann, U.: Large δ 13C Gradients in the Preindustrial North Atlantic Revealed, Science, 330, 658–659, https://doi.org/10.1126/science.1193769, 2010.
Olsen, A., Key, R. M., van Heuven, S., Lauvset, S. K., Velo, A., Lin, X., Schirnick, C., Kozyr, A., Tanhua, T., Hoppema, M., Jutterström, S., Steinfeldt, R., Jeansson, E., Ishii, M., Pérez, F. F., and Suzuki, T.: The Global Ocean Data Analysis Project version 2 (GLODAPv2) – an internally consistent data product for the world ocean, Earth Syst. Sci. Data, 8, 297–323, https://doi.org/10.5194/essd-8-297-2016, 2016.
Quay, P.: Organic Matter Export Rates and the Pathways of Nutrient Supply in the Ocean, Global Biogeochem. Cy., 37, e2023GB007855, https://doi.org/10.1029/2023GB007855, 2023.
Quay, P., Sonnerup, R., Westby, T., Stutsman, J., and McNichol, A.: Changes in the 13C/12C of dissolved inorganic carbon in the ocean as a tracer of anthropogenic CO2 uptake, Global Biogeochem. Cy., 17, https://doi.org/10.1029/2001GB001817, 2003.
Quay, P., Sonnerup, R., Stutsman, J., Maurer, J., Körtzinger, A., Padin, X. A., and Robinson, C.: Anthropogenic CO2 accumulation rates in the North Atlantic Ocean from changes in the 13C/12C of dissolved inorganic carbon, Global Biogeochem. Cy., 21, 2006GB002761, https://doi.org/10.1029/2006GB002761, 2007.
Quay, P., Sonnerup, R., Munro, D., and Sweeney, C.: Anthropogenic CO2 accumulation and uptake rates in the Pacific Ocean based on changes in the 13C/12C of dissolved inorganic carbon, Global Biogeochem. Cy., 31, 59–80, https://doi.org/10.1002/2016GB005460, 2017.
Quay, P., Emerson, S., and Palevsky, H.: Regional Pattern of the Ocean's Biological Pump Based on Geochemical Observations, Geophys. Res. Lett., 47, e2020GL088098, https://doi.org/10.1029/2020GL088098, 2020.
Quay, P. D., Tilbrook, B., and Wong, C. S.: Oceanic Uptake of Fossil Fuel CO2: Carbon-13 Evidence, Science, 256, 74–79, https://doi.org/10.1126/science.256.5053.74, 1992.
Quay, P. D., Stutsman, J., Feely, R. A., and Juranek, L. W.: Net community production rates across the subtropical and equatorial Pacific Ocean estimated from air-sea δ13C disequilibrium, Global Biogeochem. Cy., 23, 2008GB003193, https://doi.org/10.1029/2008GB003193, 2009.
Racapé, V., Pierre, C., Metzl, N., Lo Monaco, C., Reverdin, G., Olsen, A., Morin, P., Vázquez-Rodríguez, M., Ríos, A. F., and Pérez, F. F.: Anthropogenic carbon changes in the Irminger Basin (1981–2006): Coupling δ13CDIC and DIC observations, J. Mar. Syst., 126, 24–32, https://doi.org/10.1016/j.jmarsys.2012.12.005, 2013.
Rasmussen, C. E. and Williams, C. K. I.: Gaussian Processes for Machine Learning, MIT Press, Cambridge, Massachusetts, ISBN 9780262182535, https://doi.org/10.7551/mitpress/3206.001.0001, 2006.
Regier, P., Duggan, M., Myers-Pigg, A., and Ward, N.: Effects of random forest modeling decisions on biogeochemical time series predictions, Limnol. Oceanogr. Meth., 21, 40–52, https://doi.org/10.1002/lom3.10523, 2023.
Roobaert, A., Regnier, P., Landschützer, P., and Laruelle, G. G.: A novel sea surface pCO2-product for the global coastal ocean resolving trends over 1982–2020, Earth Syst. Sci. Data, 16, 421–441, https://doi.org/10.5194/essd-16-421-2024, 2024.
Samanta, S., Dalai, T. K., Pattanaik, J. K., Rai, S. K., and Mazumdar, A.: Dissolved inorganic carbon (DIC) and its δ13C in the Ganga (Hooghly) River estuary, India: Evidence of DIC generation via organic carbon degradation and carbonate dissolution, Geochim. Cosmochim. Ac., 165, 226–248, https://doi.org/10.1016/j.gca.2015.05.040, 2015.
Schmittner, A., Gruber, N., Mix, A. C., Key, R. M., Tagliabue, A., and Westberry, T. K.: Biology and air–sea gas exchange controls on the distribution of carbon isotope ratios (δ13C) in the ocean, Biogeosciences, 10, 5793–5816, https://doi.org/10.5194/bg-10-5793-2013, 2013.
Seeger, M.: Gaussian processes for machine learning, Int. J. Neural Syst., 14, 69–106, https://doi.org/10.1142/S0129065704001899, 2004.
Sharp, J. D., Fassbender, A. J., Carter, B. R., Lavin, P. D., and Sutton, A. J.: A monthly surface pCO2 product for the California Current Large Marine Ecosystem, Earth Syst. Sci. Data, 14, 2081–2108, https://doi.org/10.5194/essd-14-2081-2022, 2022.
Silverman, B. W.: Density Estimation for Statistics and Data Analysis, Monographs on Statistics and Applied Probability, Chapman and Hall, London, UK, ISBN 9780412246203, https://doi.org/10.1201/9781315140919, 1986.
Sonnerup, R. E. and Quay, P. D.: 13C constraints on ocean carbon cycle models: 13C IN OCEAN MODELS, Global Biogeochem. Cy., 26, https://doi.org/10.1029/2010GB003980, 2012.
Su, J., Cai, W.-J., Hussain, N., Brodeur, J., Chen, B., and Huang, K.: Simultaneous determination of dissolved inorganic carbon (DIC) concentration and stable isotope (δ13C-DIC) by Cavity Ring-Down Spectroscopy: Application to study carbonate dynamics in the Chesapeake Bay, Mar. Chem., 215, 103689, https://doi.org/10.1016/j.marchem.2019.103689, 2019.
Sun, Z., Li, X., Ouyang, Z., Featherstone, C., Atekwana, E. A., Hussain, N., and Cai, W.: Simultaneous onboard analysis of seawater dissolved inorganic carbon (DIC) concentration and stable isotope ratio (δ13C-DIC), Limnol. Ocean Meth., 22, 862–875, https://doi.org/10.1002/lom3.10642, 2024.
Sun, Z., Gao, H., Dong, B., Hussain, N., Atekwana, E. A., and Cai, W.: High-resolution dataset of stable carbon isotope of dissolved inorganic carbon (δ13C-DIC) from the North Atlantic Ocean, Limnol. Oceanogr. Lett., lol2.70038, https://doi.org/10.1002/lol2.70038, 2025.
Tanhua, T., van Heuven, S., Key, R. M., Velo, A., Olsen, A., and Schirnick, C.: Quality control procedures and methods of the CARINA database, Earth Syst. Sci. Data, 2, 35–49, https://doi.org/10.5194/essd-2-35-2010, 2010.
Taylor, J. R.: An introduction to error analysis: the study of uncertainties in physical measurements, 2nd Edn., University Science Books, Sausalito, Calif, 327 pp., ISBN 9780935702750, 1997.
Wanninkhof, R., Doney, S. C., Bullister, J. L., Levine, N. M., Warner, M., and Gruber, N.: Detecting anthropogenic CO2 changes in the interior Atlantic Ocean between 1989 and 2005, J. Geophys. Res., 115, 2010JC006251, https://doi.org/10.1029/2010JC006251, 2010.
Wu, Z., Lu, W., Roobaert, A., Song, L., Yan, X.-H., and Cai, W.-J.: A machine-learning reconstruction of sea surface pCO2 in the North American Atlantic Coastal Ocean Margin from 1993 to 2021, Earth Syst. Sci. Data, 17, 43–63, https://doi.org/10.5194/essd-17-43-2025, 2025.
Yang, B., Emerson, S. R., and Quay, P. D.: The Subtropical Ocean's Biological Carbon Pump Determined From O2 and DIC/DI 13C Tracers, Geophys. Res. Lett., 46, 5361–5368, https://doi.org/10.1029/2018GL081239, 2019.
- Abstract
- Introduction
- Data and Methods
- Results and Discussion
- Data availability
- Code availability
- Conclusions
- Appendix A: Numerical Model-based Validation of the GPR Method for 4D δ13CDIC Reconstruction
- Author contributions
- Competing interests
- Disclaimer
- Acknowledgements
- Financial support
- Review statement
- References
- Abstract
- Introduction
- Data and Methods
- Results and Discussion
- Data availability
- Code availability
- Conclusions
- Appendix A: Numerical Model-based Validation of the GPR Method for 4D δ13CDIC Reconstruction
- Author contributions
- Competing interests
- Disclaimer
- Acknowledgements
- Financial support
- Review statement
- References