the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
National-scale sub-meter mapping of Spartina alterniflora in mainland China 2020
Bingfeng Zhou
Meng Xu
Mingming Jia
Dehua Mao
Kai Cheng
Xiumin Zhu
Haoyue Jiang
Jie Song
Yinghai Ke
Zhenxin Zhang
Yue Huang
Miaojing Wei
Xiaojuan Li
Huili Gong
Current large-scale maps of Spartina alterniflora (S. alterniflora) with 10 m resolution hinder accurate delineation of community boundaries, detection of internal features such as creeks, and identification of small patches. These limitations further compromise the accuracy of spatial distribution extraction and subsequent analyzes. To this end, this study produced the first 2020 national-scale Sub-meter (0.9 m) S. alterniflora Map of Mainland China (CM-SSM), using an object- and sub-meter-enhanced pixel-based phenological feature composite method. The method integrates phenological features from Sentinel-2 with spatial and texture details from Google Earth imagery, improving the spectral separability and mitigating mixed-pixel effects. Compared to the 10 m S. alterniflora product of Mainland China (CMSA), CM-SSM improved overall accuracy by 14.60 % and the F1 score by 0.24. Although the total mapped areas of CM-SSM (59 371 ha) and CMSA (58 006 ha) differ by only 1365 ha, their spatial distributions diverge substantially. When benchmarked against CM-SSM, CMSA exhibited commission and omission errors totaling 34 273 ha (57.73 %). Moreover, the number of patches identified by CM-SSM (148 072) was over 17 times greater than that of CMSA, reflecting its superior capability in detecting fragmented distributions. In addition, Soil Organic Carbon (SOC) estimates derived from CM-SSM were 706.69 Gg (23.09 %) higher than those reported by the corresponding national SOC product for the same year, emphasizing the essential contribution of high-resolution mapping to accurate carbon accounting for S. alterniflora. These advances enhance understanding of S. alterniflora invasion dynamics, support carbon accounting, and inform evidence-based coastal wetland management and restoration. The map is available at https://doi.org/10.5281/zenodo.16296823 (Xu et al., 2025).
- Article
(15774 KB) - Full-text XML
-
Supplement
(672 KB) - BibTeX
- EndNote
Spartina alterniflora (S. alterniflora), native to the Atlantic coast of North America and widely recognized as a classic “ecosystem engineer”, was intentionally introduced into China in 1979 to enhance embankment stability and mitigate coastal erosion (Jackson et al., 2021; Liu et al., 2020). Due to its lack of natural predators and high reproductive capacity, S. alterniflora has rapidly expanded along China's coast over the past four decades, reaching a total extent of more than 50 000 ha (Meng et al., 2020). This rapid spread has posed serious threats to coastal ecosystems, including the displacement of native species, degradation of nearshore habitats, and measurable declines in biodiversity (Li et al., 2009; Okoye et al., 2020). In response, a range of control strategies, including physical removal, chemical control, and biological replacement, were employed to manage the invasion of S. alterniflora in China (Zheng et al., 2023a). However, the effectiveness of these measures varied significantly across regions, and the risk of reinvasion remained high (Li et al., 2022a; Zhao et al., 2020). Given the rapid natural spread of S. alterniflora and the continued influence of human activities on its spatial distribution, there is an urgent need for accurate, large-scale monitoring to delineate its distribution patterns and assess the effectiveness of removal effort.
Remote sensing has been widely used for mapping S. alterniflora due to its capability for large-area coverage and long-term, repeatable monitoring (Chen et al., 2020; Lourenço et al., 2021; Lv et al., 2019). Existing mapping products can be divided into two categories based on spatial resolution. The first category includes products derived from High Resolution (HR) imagery (10–30 m), such as Landsat and Sentinel series data (Zuo et al., 2012, 2025). Liu et al. (2018) produced the S. alterniflora map along mainland China's coast in 2015 at 30 m using Landsat 8. Subsequently, Hu et al. (2021) generated a 2019 coastal saltmarsh map (including S. alterniflora) using Sentinel-1, with an improved spatial resolution from 30 to 10 m. To analyze expansion dynamics of S. alterniflora, Zhang et al. (2017) mapped a sparse multi-temporal dataset spanning 1990 to 2014 using Landsat time-series imagery. Similarly, Mao et al. (2019) produced S. alterniflora maps at 5- or 10-year intervals from 1990 to 2015. The mapping intervals in previous studies were typically greater than 5 years, making it difficult to capture the ongoing spread of S. alterniflora. Therefore, Li et al. (2024) generated annual distribution maps of S. alterniflora between 2017 and 2021 using Sentinel-2 imagery, enabling precise monitoring of interannual changes.
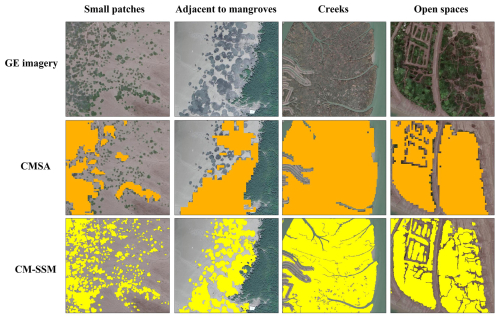
Nevertheless, relying solely on HR imagery (10–30 m) presents three key challenges due to inherent spatial limitations. First, existing products often fail to detect small patches of S. alterniflora, which are ecologically important and may function as early indicators of invasion risk (Chen et al., 2020). Second, boundaries between S. alterniflora and co-occurring species remain poorly defined, limiting accurate delineation of its spatial extent. For instance, in mangrove areas, boundary misclassification hinders reliable assessment of invasion risk (Zheng et al., 2023b). Third, internal features within S. alterniflora communities, such as creeks and open spaces, are difficult to capture. Creeks, in particular, influence hydrological processes and seed dispersal, thereby affecting the rate and extent of spread (Sun et al., 2020). These issues obstruct the effective application of existing products in precise monitoring and ecological management.
The second category includes products derived from Very High Resolution (VHR) imagery (finer than 10 m), such as UAV (Windle et al., 2023), WorldView (Dong et al., 2024), Gaofen (Li et al., 2021), and SPOT imagery (Liu et al., 2017a). Compared with HR data, these sources offer improved spatial detail and effectively reduce classification errors caused by mixed pixels. However, their high acquisition cost and limited spatial coverage constrain their use in large-scale applications. The Google Earth (GE) platform provides free access to sub-meter imagery with rich spatial detail (Li et al., 2022b), creating new opportunities for large-area, high-precision monitoring of S. alterniflora. Nevertheless, the lack of multispectral information in GE imagery restricts its utility for spectral-based identification (Zhou et al., 2024).
Phenology-based methods, which exploit spectral variations during vegetation growth, are widely recognized for their effectiveness in mapping S. alterniflora (Zeng et al., 2022; Zhang et al., 2022). Initially, single-date imagery from the growing period of S. alterniflora was used for classification (Ouyang et al., 2013; Wang et al., 2015a). However, relying solely on the growing period is insufficient for accurate mapping because S. alterniflora shares similar spectral features with other saltmarsh vegetation, especially evergreen mangroves (Ai et al., 2017; Sun et al., 2021). To address this, time-series imagery has been used to construct phenological trajectories, enabling improved distinction through the integration of multiple growth phases (Sun et al., 2016; Liu et al., 2017b). For example, Sun et al. (2016) constructed monthly NDVI time-series from the Chinese HuanJing-1 satellite imagery to monitor salt marsh vegetation, including S. alterniflora. While this method shows strong potential, two major challenges remain. Frequent cloud cover and tidal disturbances in coastal regions complicate the acquisition of high-quality, high-temporal-resolution imagery. Moreover, S. alterniflora exhibits spatial phenological heterogeneity, where communities in different regions may be at different phenological stages simultaneously, introducing spectral inconsistencies that reduce classification stability and accuracy.
To address the limitations mentioned above, Tian et al. (2020a) proposed a Pixel-based Phenological Feature (PPF) composite method. This method leverages the computational capacity and extensive data resources of the Google Earth Engine (GEE) platform to perform image compositing at the pixel level, aiming to overcome the difficulty of acquiring high-quality imagery in intertidal zones. Instead of relying on entirely cloud-free scenes, the method integrates all cloud-free pixels across multiple images, significantly improving the utilization of available data and alleviating the scarcity of usable imagery. Two key phenological periods (green period and senescence period) were selected based on their spectral distinctiveness. During the green period, S. alterniflora is spectrally distinguishable from non-vegetated surfaces such as mudflats and water; in the senescence period, it is more separable from evergreen species like mangroves. These periods provide complementary spectral features, enhancing the separability of S. alterniflora and capturing most of its relevant phenological information, while minimizing interference from transitional periods. Furthermore, to account for the phenological variability of S. alterniflora across different geographic locations, the method constructs composite imagery by extracting the greenest pixels during the green period and the most senescent pixels during the senescence period. This selection of extreme phenological states effectively reduces spatial heterogeneity in the phenology of S. alterniflora. Subsequent studies have confirmed the value of the dual-temporal phenological feature composite method. It has been shown to improve S. alterniflora classification accuracy (Zhang et al., 2020, 2023b) and to perform well in broader coastal wetland mapping (Chen and Kirwan, 2022; Zhao et al., 2023), demonstrating its effectiveness and reliability. However, these studies relied on pixel-based classification methods, which consider only the spectral value of individual pixels and ignore spatial relationships with neighboring pixels, thus limiting classification accuracy (Frohn et al., 2011). Independent pixel-wise classification often results in isolated misclassified pixels, producing the salt-and-pepper effect (Dronova, 2015). In contrast, object-based image analysis (OBIA) integrates shape, texture, and spatial context features and takes advantage of the spectral consistency within image objects. This method has shown promising potential for S. alterniflora identification (Wang et al., 2021). Nevertheless, the application of OBIA in large-scale, sub-meter S. alterniflora mapping using VHR imagery remains underexplored.
This study aims to establish an object-based, large-scale mapping approach by integrating multi-source remote sensing imagery to produce the first sub-meter map of S. alterniflora across mainland China. Our objectives are threefold: (1) to develop a novel Object- and Sub-meter-enhanced PPF (OSPPF), (2) to produce a 2020 Sub-meter S. alterniflora Map of Mainland China (CM-SSM), and (3) to evaluate the significant improvements of CM-SSM over the latest 10 m resolution map of S. alterniflora. Overall, the OSPPF effectively mitigates the mixed-pixel problem by incorporating sub-meter GE imagery. Additionally, by integrating OBIA, the proposed OSPPF approach leverages multi-dimensional features such as texture, shape, and spatial context, thereby overcoming the limitations of pixel-based classification that relies solely on spectral features. This integration enhances the classification accuracy of S. alterniflora. CM-SSM addresses issues in existing products, such as inaccurate boundary depiction of S. alterniflora, poor identification of internal details, and limited ability to detect small patches. As a high-resolution and high-accuracy map, CM-SSM provides a robust data foundation for management assessment, blue carbon stock estimation, and coastal sustainable development.
2.1 Study area
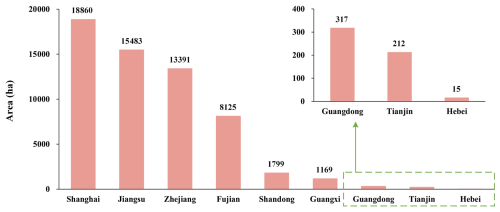
S. alterniflora is distributed across nine provinces in mainland China (20–41° N, 108–122° E), with coverage areas ranked in descending order as follows: Shanghai, Jiangsu, Zhejiang, Fujian, Shandong, Guangxi, Guangdong, Tianjin, and Hebei. Common co-occurring species include Phragmites australis, Suaeda salsa, Tamarix chinensis, and mangroves.
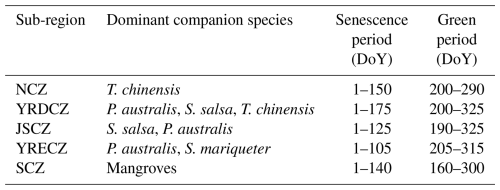
To preliminarily delineate the potential distribution of S. alterniflora, a 10 km coastal buffer zone was generated by extending seaward from the coastline dataset. Considering the regional differences in co-occurring species and phenology, the coastal buffer was subdivided into five subregions: the Southern Coastal Zone (SCZ), Yangtze River Estuary Coastal Zone (YRECZ), Jiangsu Coastal Zone (JSCZ), Yellow River Delta Coastal Zone (YRDCZ), and Northern Coastal Zone (NCZ). Subsequently, two national-scale S. alterniflora products from 2020 (see Sect. 2.2.2) were collected to extract their union, which was then expanded with a 100 m buffer to cover potential edge areas. Omission errors were manually corrected through visual interpretation of VHR imagery. The red-highlighted area represented an S. alterniflora patch. The study area was divided into five subregions, each containing multiple S. alterniflora patches. All Sentinel-2 and GE imagery were collected within these subregions (Fig. 1).
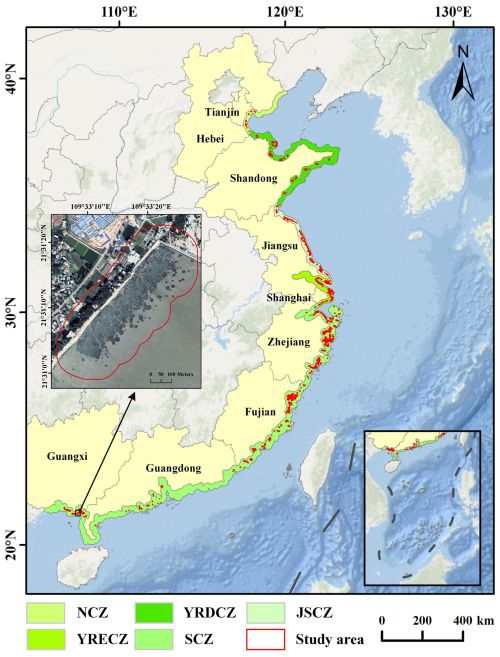
Figure 1Location of the study site in the coastal zone of mainland China. The background imagery is provided by Esri (https://www.esri.com, last access: 23 July 2025) and its data partners. The VHR imagery in the figure is from © Google Earth 2020.
2.2 Datasets
2.2.1 Remote sensing imagery
This study acquired approximately 6007 Sentinel-2 Surface Reflectance (SR) images from the year 2020 through the GEE platform. As S. alterniflora grows in intertidal zones and is highly susceptible to cloud contamination and tidal variation, both scene-based and pixel-based methods were applied to ensure image quality (Chen et al., 2025). At the scene level, images with more than 70 % cloud cover within the study area were excluded based on metadata attributes (Ni et al., 2021). At the pixel level, bitwise operations were used to the Sentinel-2 Scene Classification Layer (SCL) to mask cloud (SCL = 7–9), cirrus (SCL = 10), and cloud shadows (SCL = 3). To further reduce tidal effects on classification, water pixels (SCL = 6) were also removed. Additionally, 0.9 m GE imagery from 2020 with RGB bands was selected under low-tide and cloud-free conditions. In regions lacking high-quantity 2020 imagery, supplementary GE imagery from 2019 or 2021 was used.
2.2.2 Existing S. alterniflora products
Two S. alterniflora products covering coastal mainland China in 2020 were collected for study area delineation and comparative analysis (Table 1). Mao et al. (2019) developed a multi-temporal S. alterniflora dataset (1990–2015) and used it to generate a 30 m resolution product in 2020 (hereafter referred to as SpProduct_30m). More recently, Li et al. (2024) generated a 10 m resolution S. alterniflora map of mainland China for 2020 (CMSA) using Sentinel-2 imagery. Both SpProduct_30m and CMSA served as critical references for delineating the study area (see Sect. 2.1). As the highest-resolution national-scale S. alterniflora product for 2020, CMSA was used as the benchmark. Specifically, we evaluated the improvement in detection capability achieved by our sub-meter product relative to the CMSA.
2.2.3 Reference data
The ecological complexity of intertidal zones where S. alterniflora grows poses challenges for large-scale field sampling and limits the availability of sufficient reference data. To address this issue, we constructed a high-quality sample dataset by integrating field surveys with multi-source VHR imagery. First, field surveys were conducted in 2020 in typical S. alterniflora habitats, including the Beibu Gulf, Jiulong River Estuary, and Zhangjiang Estuary. Using differential GPS, we collected 1396 ground validation points, including 661 S. alterniflora and 735 non-S. alterniflora samples. Subsequently, VHR imagery temporally aligned with the field survey was collected from UAVs, the Gaofen series, and GE. Based on this, experienced researchers visually interpreted and manually labeled 10 570 S. alterniflora and 20 247 non-S. alterniflora points by overlaying field data with VHR imagery. The non-S. alterniflora class includes co-occurring species (e.g., mangroves, Phragmites australis, Suaeda salsa) and other land covers such as mudflats. All sample points were evenly distributed across the five subregions defined in Sect. 2.1. Finally, a stratified random sampling method was used to divide S. alterniflora and non-S. alterniflora sample points within each subregion into training and validation sets at a 7:3 ratio. In addition, the class ratio of S. alterniflora to non-S. alterniflora (approximately 1:2) was preserved during the splitting process. The sample points from all subregions were then merged to form a complete reference dataset. This method mitigated issues of spatial autocorrelation and class imbalance (Wang et al., 2020). The training set was used to select input features for the Random Forest (RF) classifier (see Sect. 2.3.3), while the validation set was employed to assess classification accuracy.
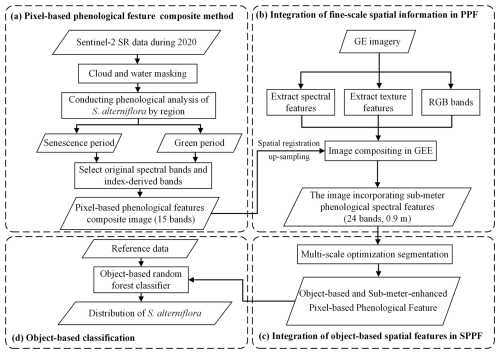
2.3 Development of an object- and sub-meter-enhanced PPF
This study proposed an Object- and Sub-meter-enhanced Pixel-based Phenological Feature (OSPPF) composite method for mapping S. alterniflora, including four steps (Fig. 2). First, a Pixel-based Phenological Feature (PPF) was constructed using Sentinel-2 imagery (10 m). Second, spatial and texture features extracted from GE imagery (0.9 m) were integrated to enhance the PPF, resulting in the Sub-meter-enhanced PPF (SPPF). Third, a multi-scale object-based segmentation strategy was used to extract the OSPPF. Finally, a RF classifier was applied to generate the initial result, which was then manually refined to generate the final S. alterniflora distribution map.
2.3.1 Pixel-based phenological feature composite method
Phenological features are critical for identifying S. alterniflora. Previous studies have shown that the green and senescence periods are two key phenological phases that enhance the spectral separability of S. alterniflora from background land covers (Tian et al., 2020a). Accordingly, this study constructed annual NDVI time series curves based on Sentinel-2 imagery acquired between 1 January and 31 December 2020, to determine the green and senescence periods for each subregion. Using JSCZ as an example, 175 pure S. alterniflora pixels were selected through visual interpretation of GE imagery, ensuring an even spatial distribution. NDVI values for these pixels were calculated from cloud-masked Sentinel-2 imagery, extracting the median value for each Day of Year (DoY) to construct the NDVI time series. To reduce noise caused by cloud cover and atmospheric effects, the NDVI time series was smoothed using the Savitzky–Golay (SG) filter (Savitzky and Golay, 1964). As shown in Fig. 3, the NDVI time series of JSCZ exhibited a phenological pattern consistent with that reported by Tian et al. (2020a). To determine the two key phenological periods of S. alterniflora in the JSCZ, the annual NDVI frequency distribution histogram and its first derivative curve were generated based on 175 pure S. alterniflora pixels (Fig. S1 in the Supplement). As shown in Fig. S1, the NDVI values exhibited a marked decline around 0.3 and a sharp increase around 0.5, corresponding to the transitions from the senescence to the transitional period and from the transitional to the green period, respectively. Therefore, NDVI values below 0.3 during DoY 1–125 indicated the senescence period, whereas values above 0.5 during DoY 190–325 corresponded to the green period. Following this procedure, the key phenological windows were identified for each subregion (Table 2).
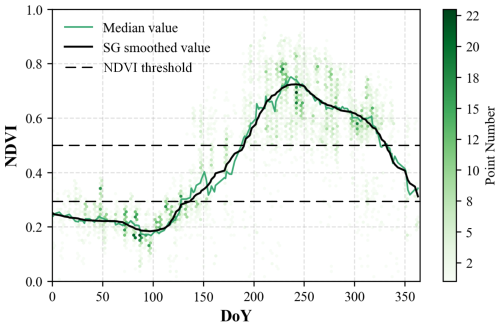
Figure 3NDVI time series analysis of the S. alterniflora in JSCZ. Point density is represented using hexagonal binning, with color intensity indicating the concentration of data points.
Based on the two identified phenological periods, the PPF was constructed by integrating vegetation indices and original spectral bands. Specifically, five indices were selected to characterize the phenological periods (Table 3): Normalized Difference Vegetation Index (NDVI), Enhanced Vegetation Index (EVI), Plant Senescence Reflectance Index (PSRI), Normalized Difference Water Index (NDWI), and Land Surface Water Index (LSWI). NDVI, EVI, and NDWI were used during the green period, while PSRI and LSWI characterized the senescence period. The selection rationale is as follows: (1) NDVI is sensitive to green vegetation but tends to saturate under dense canopies, whereas EVI remains responsive at high biomass levels, making them complementary during the green period (Ni et al., 2021). (2) NDWI captures reflectance differences between vegetation and water in the green and near-infrared bands, effectively distinguishing S. alterniflora from water when canopy cover is high during the green period (Mancino et al., 2020). (3) PSRI responds to changes in carotenoid pigments associated with senescence, making it suitable for detecting vegetation during the senescence period (Tian et al., 2020a). (4) As LSWI is sensitive to leaf water content, the progressive moisture loss in S. alterniflora during senescence leads to decreased LSWI values, improving its separability from moist backgrounds such as mudflats and water (Wu et al., 2020). Each index was derived from Sentinel-2 imagery of its corresponding phenological period, with median values of valid observations computed per pixel to generate the composite images.
In addition, five original bands of Sentinel-2 were selected for both phenological periods: B2 (blue), B3 (green), B4 (red), B8 (NIR) and B11 (SWIR 1). B2, B3, and B4 cover the visible spectrum and are useful for distinguishing S. alterniflora from water and mudflats. B8 and B11 are sensitive to vegetation structure and moisture content, effectively capturing spectral transitions of S. alterniflora from the green to senescence period. Finally, the vegetation indices and selected spectral bands for both phenological periods (a total of 15 bands) were integrated to construct the PPF composite images.
2.3.2 Integration of fine-scale spatial information in PPF
To address the spatial resolution limitations of PPF, this study introduced the SPPF composite method, incorporating fine-scale spatial information from GE imagery through three steps: GE feature extraction, spatial-scale normalization and geometric registration, and image compositing.
First, spectral and texture features were extracted from GE imagery. For spectral features, the Normalized Green-Blue Difference Index (NGBDI) and the Normalized Green-Red Difference Index (NGRDI), derived from the RGB bands (Table 4), have proven effective for wetland vegetation classification (Zheng et al., 2022). Texture features were computed from the red band using the Grey-Level Co-Occurrence Matrix (GLCM) method (Haralick et al., 2007), extracting four second-order statistics commonly used in vegetation classification: contrast, entropy, correlation, and homogeneity (Wang et al., 2015b). Sliding window size is critical to texture extraction, with small windows failing to capture spatial texture and large ones blurring object boundaries. A 17×17 sliding window was applied to the grayscale image of the red band (Li et al., 2020), generating GLCMs and computing four texture metrics per window to produce corresponding texture bands.
Second, enabling effective integration of multi-source data required resampling to a common resolution and geometric registration. The Sentinel-2 spectral bands and associated vegetation index images (10–20 m) were resampled using cubic convolution to match the 0.9 m GE imagery. Then, GE imagery was used as the reference to selecting evenly distributed and clearly identifiable control points from both image sources (e.g., tidal creek intersections, aquaculture pond corners, and vegetation patch boundaries). These points were used to construct a polynomial transformation model for registering the Sentinel-2 imagery. Finally, phenological features derived from Sentinel-2 were integrated with the spectral, texture, and RGB features extracted from GE imagery to construct the SPPF composite images consisting of 24 bands.
2.3.3 Integration of object-based spatial features in SPPF
Considering the complex boundaries and homogeneous interiors of S. alterniflora patches, accurately delineating their edges remains challenging when using pixel-based features. Therefore, we developed an object-based feature extraction method that incorporated a multi-scale optimized segmentation strategy, enabling the effective integration of spatial context and pixel neighborhood relationships for improved boundary detection. The method included two key steps: identifying the boundary regions of S. alterniflora patches and determining the optimal multi-scale segmentation parameters.
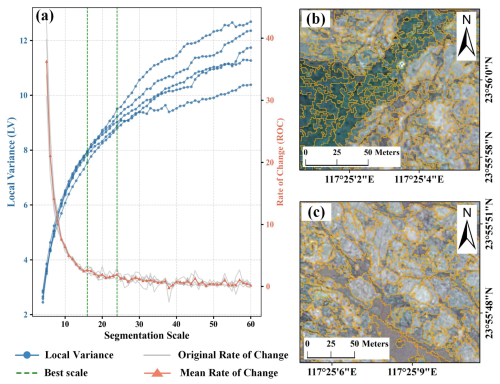
Figure 4(a) Variation of Local Variance (LV) and Rate of Change (ROC) with Scale Parameter. (b) Segmentation results with scale parameter of 16. (c) Segmentation results with scale parameter of 24. The VHR imagery in the figure is from © Google Earth 2020.
To delineate patch boundaries, each patch identified from the CMSA was expanded outward and contracted inward by 10 m, corresponding to the spatial resolution of the CMSA. This process resulted in a 20 m annular buffer zone that captures the complex transitional areas along the edges of S. alterniflora patches while minimizing the inclusion of non-target features, thereby ensuring both the accuracy and representativeness of boundary identification. To implement the multi-scale optimized segmentation strategy, the Estimation of Scale Parameter (ESP) method was applied to identify the optimal scales for both edge-complex and interior-homogeneous regions. ESP quantifies image region homogeneity by computing Local Variance (LV) and its Rate of Change (ROC) across multiple scales, with ROC peaks typically indicating optimal segmentation (Drăguţ et al., 2010). To improve scale representativeness, one typical region was selected from each of the five subregions (see Sect. 2.1), and GE imagery was segmented to calculate LV and ROC. Following Wang et al. (2021), the scale range was set to 4–60 with a step of 1. The shape and compactness parameters were set to 0.1 and 0.5, respectively (Wan et al., 2014). As shown in Fig. 4a, the mean ROC curve across the five regions exhibited multiple peaks, indicating several candidate optimal scales. Based on these peaks, a series of segmentation results were visually interpreted. Scale 16 was optimal for capturing fine details in boundary-complex regions, while scale 24 was better represented the homogeneous interior, balancing spatial coherence with processing efficiency. Segmentation results for both scales are shown in Fig. 4b and c. Based on the determined scale parameters, object-based segmentation was performed on the SPPF composite imagery in eCognition, producing the OSPPF for subsequent classification.
2.3.4 Classifier selection and parameterization
Although deep learning has shown potential for sub-meter mapping of S. alterniflora at the local scale, its reliance on high-quality training samples and the limited generalization ability of models constrain its application at the national scale (Zhou et al., 2024). In contrast, the object-based RF classifier integrates spectral, textural, and spatial contextual features, demonstrating higher stability and classification accuracy in identifying S. alterniflora. Moreover, it outperforms the pixel-based RF method in mapping S. alterniflora (Tian et al., 2020b; Yan et al., 2021). Therefore, we used the object-based RF classifier in eCognition. The RF algorithm aggregates predictions from multiple decision trees, with the number of trees being a key parameter influencing classification performance (Breiman et al., 2001). To determine the optimal number of trees, a sensitivity analysis was conducted using the training and validation datasets (see Sect. 2.2.3), varying the tree count from 50 to 500 at intervals of 50. The results indicated that 200 trees yielded the highest overall accuracy on the validation set. The multi-source features and training samples were then input into the object-based RF classifier to produce the Initial Sub-meter S. alterniflora Map of Mainland China (ICM-SSM). To enhance accuracy, experienced researchers visually interpreted GE imagery and corrected the ICM-SSM. Consequently, the final Sub-meter S. alterniflora Map of Mainland China (CM-SSM) was generated.
2.4 Accuracy assessment
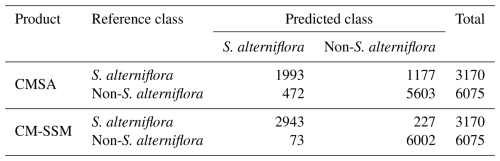
To assess the mapping effectiveness of the OSPPF method for S. alterniflora, two sets of comparative experiments were designed in typical area to generate comparative results. Each result was compared with the CM-SSM generated by OSPPF, focusing on boundary delineation, small patch detection, and internal structure extraction. Classification accuracy of the CM-SSM was quantitatively assessed using confusion matrix-based metrics. Producer accuracy (PA), user accuracy (UA), overall accuracy (OA), and the F1 score were calculated using the validation dataset, which comprised 3170 positive and 6075 negative samples (see Sect. 2.2.3). To further evaluate the performance of CM-SSM, a comparative analysis was conducted against the CMSA. The comparison included both classification detail and overall statistics. At the detail level, attention was paid to differences in edge, internal structure, and small patch. At the statistical level, we quantified differences in total area, number of patches, and spatial distribution of S. alterniflora.
3.1 Performance of OSPPF
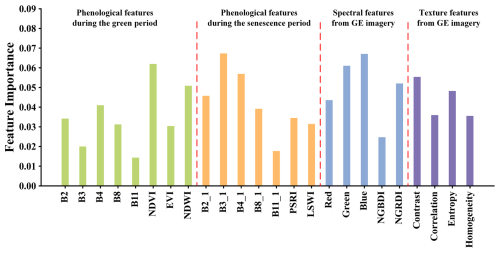
To assess the contribution of GE imagery to classification performance, S. alterniflora mapping was conducted using two methods in the Dandou Sea: (1) object-based PPF (OPPF) classification using resampled Sentinel-2 imagery alone, and (2) OSPPF classification integrating both Sentinel-2 and GE imagery. As shown in Fig. 6a, classification based solely on Sentinel-2 imagery was able to capture the general outline of S. alterniflora communities but failed to effectively delineate open spaces within the patches. In addition, Fig. 6b demonstrates that small, fragmented S. alterniflora patches were poorly detected, and the boundaries between S. alterniflora and mangroves were inaccurately represented. In contrast, the CM-SSM generated using fused GE imagery exhibited superior spatial detail, successfully identifying small patches and internal details, as well as accurately delineating boundaries between S. alterniflora and co-occurring species. Furthermore, we conducted a feature importance analysis using the RF classifier (Zhang et al., 2023a). As shown in Fig. 5, spectral and texture features derived from GE imagery consistently contributed highly to the classification of S. alterniflora. This can be primarily attributed to the rich spatial texture information provided by GE imagery, which effectively complements the phenological features and thereby enhances classification accuracy.
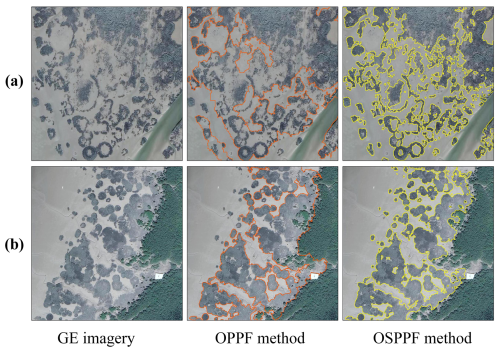

Figure 6Comparison of classification results using OPPF and OSPPF methods in Dandou Sea. The result generated by the OSPPF method is the final, manually refined CM-SSM product. The VHR imagery in the figure is from © Google Earth 2020.
To further compare pixel-based and object-based classification methods in S. alterniflora mapping, the SPPF and OSPPF methods were applied in the Dandou Sea. As illustrated in Fig. 7, the CM-SSM generated using object-based classification accurately extracted the boundaries between S. alterniflora and surrounding land cover types such as mudflats and mangroves, whereas the pixel-based method showed poor boundary delineation. Moreover, Fig. 7 indicates that the pixel-based result suffered from salt-and-pepper noise, particularly within and along the edges of S. alterniflora patches. In contrast, the CM-SSM demonstrated a smoother spatial distribution and effectively suppressed such noise.
3.2 Comparison with the latest 10 m product
3.2.1 Accuracy assessment
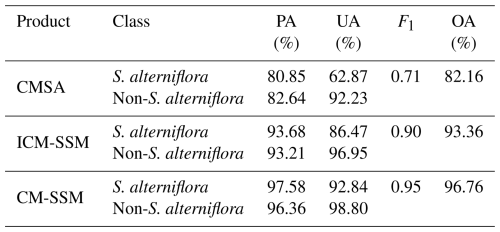
Table 5 presents the accuracy assessment of three products (CMSA, ICM-SSM, and CM-SSM) based on validation samples (see Sect. 2.2.3). Among them, CM-SSM achieved the best classification performance, with OA and F1 scores of 96.76 % and 0.95, respectively. ICM-SSM also performed well, with an OA of 93.36 % and an F1 score of 0.90, although slightly lower than those of CM-SSM. This difference is mainly attributed to the manual refinement, which improved boundary delineation and the identification of small patches. In contrast, CMSA exhibited the lowest accuracy, with OA and F1 scores 14.60 % and 0.24 lower than those of CM-SSM, respectively. Given the wide latitudinal span of the study area, the spatial variability in classification performance was further examined by evaluating CMSA and CM-SSM across five subregions using validation samples. The results show that CM-SSM consistently achieved superior performance, with OA exceeding 95.00 % and F1 scores above 0.90 in all subregions (Tables S1–S5 in the Supplement). To further assess whether the accuracy improvement of CM-SSM over CMSA is statistically significant, we applied McNemar's test based on validation samples. The results indicated a statistically significant difference (χ2=820.22, p<0.05), confirming the robustness of the improvements in OA and F1 scores (McNemar, 1947). The superior performance of CM-SSM is mainly due to its reduction in both omission and commission errors, which contributed to the higher OA and F1 scores, as shown in Table 6.
3.2.2 Spatial details
To reveal the difference between CM-SSM and CMSA, a comparative analysis focusing on small patches, boundaries, and internal structural features of S. alterniflora communities was conducted (Fig. 8). First, given the fragmented distribution of S. alterniflora, numerous small patches are typically present. CM-SSM effectively detected nearly all of them, whereas CMSA missed most. Second, S. alterniflora often encroaches upon mangrove habitats due to its aggressive spread, increasing boundary complexity. CM-SSM showed superior delineation of the interface between the two species, clearly depicting invasion fronts that CMSA did not capture effectively. Finally, intertidal S. alterniflora communities often include internal features like creeks and open spaces. CM-SSM successfully resolved open spaces and narrow creeks, while CMSA lacked the spatial detail to do so.
3.2.3 Spatial distribution statistics
A total of 148 072 patches were identified in CM-SSM, approximately 17 times more than CMSA. The total mapped area of CM-SSM reached 59 371 ha, exceeding that of CMSA by 1365 ha. To further compare patch size and area differences between the two products, statistics were summarized across six area classes defined by the minimum mapping unit of CMSA (i.e., one 10 m pixel). These intervals included: 0.01 ha (1 pixel), 0.1 ha (10 pixels), 1 ha (100 pixels), 100 ha (10 000 pixels), 1000 ha (100 000 pixels), and greater than 1000 ha (Fig. 9). In addition, to assess spatial distribution differences, an overlay analysis was conducted using CM-SSM as the reference (Figs. 10 and 11).
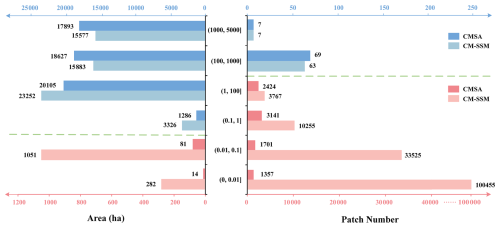
Figure 9Zonal statistics of the number and area of S. alterniflora patches identified by CM-SSM and CMSA.
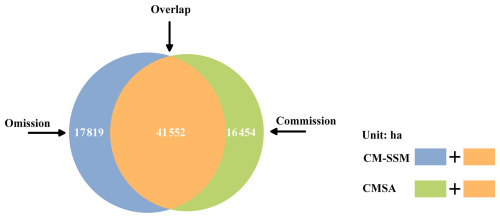
Figure 10Spatial distribution difference statistics of S. alterniflora identified by CM-SSM and CMSA.
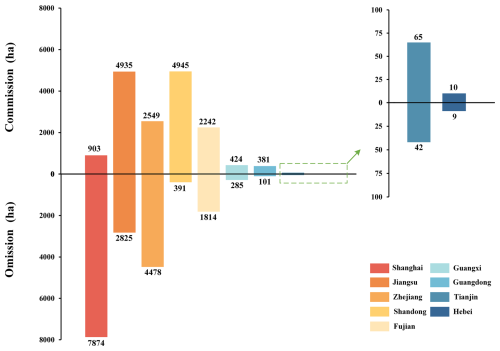
Figure 11Using CM-SSM as the reference, the commission and omission of S. alterniflora area by province were calculated for CMSA.
As shown in Fig. 9, when patch area was no greater than 0.1 ha, CM-SSM detected 44 times more patches than CMSA, with a total area 14 times larger. This notable difference underscores CM-SSM's superior ability to capture small and fragmented S. alterniflora communities. In CM-SSM, such small patches accounted for 90.48 % of the total patch count but only 2.25 % of the total area. CMSA reported 35.15 % and 0.16 % for the same class. These results are consistent with the highly fragmented spatial distribution of S. alterniflora. In terms of large patches exceeding 100 ha, both products demonstrated similar performance. In CM-SSM, these patches represented just 0.05 % of the total number but contributed 52.99 % of the total area, while in CMSA the corresponding proportions were 0.87 % and 62.96 %.
When benchmarked against CM-SSM, CMSA exhibited 16 454 ha of commission and 17 819 ha of omission, resulting in a total spatial mismatch of 34 273 ha (Fig. 10). Notably, the overall area difference between the two products was small because the commission and omission nearly offset each other. To further explore spatial discrepancies, a province-level analysis was conducted. As shown in Fig. 11, both the pattern and magnitude of these differences varied across provinces. For example, in Shanghai, omission was the primary contributor to spatial disagreement, whereas in Jiangsu, commission and omission were more balanced, together accounting for 7760 ha difference.
3.3 Mapping results of mainland China
Figure 12a illustrates the 2020 distribution of S. alterniflora across mainland China based on CM-SSM. The results indicate that S. alterniflora was densely distributed along the coasts of Fujian, Zhejiang, Shanghai, and Jiangsu, where nearly all large-scale communities were located. These four provinces also reported the largest total areas, together accounting for 94.08 % of the total in mainland China (Fig. 13). In Guangxi, large patches were primarily distributed around the Beibu Gulf, while in Guangdong, S. alterniflora appeared more scattered and patchier. In northern provinces such as Shandong, Hebei, and Tianjin, S. alterniflora communities were generally small in size and spatially fragmented. To better illustrate spatial patterns across ecosystems, three representative regions were selected. As shown in Fig. 12b and d, Dandou Sea and the Zhangjiang Estuary reflect zones of S. alterniflora-mangrove coexistence, where S. alterniflora is gradually encroaching into mangrove habitats. In contrast, Fig. 12c highlights northern Jiangsu's mudflats, where S. alterniflora has colonized unvegetated mudflats and formed expansive stands, demonstrating strong invasiveness and ecological adaptability.
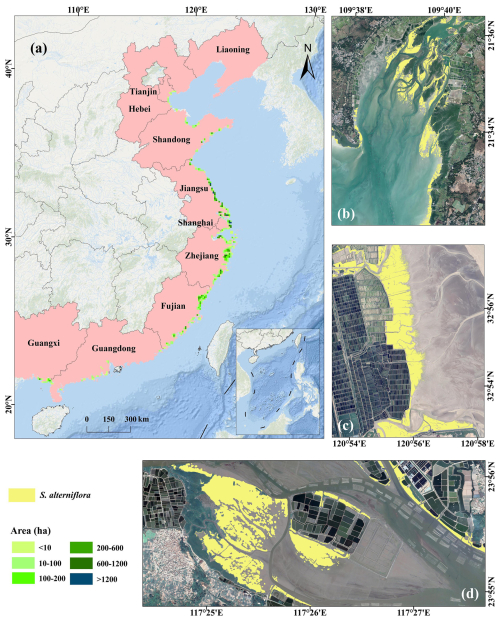
Figure 12(a) Spatial distribution of S. alterniflora in mainland China for 2020, derived from the CM-SSM, with each grid cell representing the total S. alterniflora area within a 10 km × 10 km unit; (b–d) CM-SSM distribution map in typical areas, including (b) Dandou Sea, (c) mudflats of northern Jiangsu, and (d) Zhangjiang Estuary. The background imagery is provided by Esri (https://www.esri.com, last access: 23 July 2025) and its data partners. The VHR imagery in the figure is from © Google Earth 2020.
4.1 The advantages of the OSPPF
This study proposed a novel OSPPF composite method that enhanced the PPF by integrating GE imagery and object-based classification. Previous studies have demonstrated that PPF improves the spectral separability of S. alterniflora by utilizing dual-phase phenological information (Tian et al., 2020a; Li et al., 2024). However, PPF based solely on Sentinel-2 imagery is limited in capturing spatial detail due to its 10 m resolution, despite providing rich phenological information. To overcome this limitation, GE imagery was integrated to enhance spatial detail (Fig. 6), offering two key advantages. First, GE imagery mitigates the mixed-pixel problem. Patch-level statistics show that 100 455 patches are smaller than 100 m2, accounting for 67.84 % of all patches (Fig. 9). These patches are often overlooked in Sentinel-2 classifications due to mixed-pixel problem, yet they are critical indicators of early-stage invasion. In contrast, the sub-meter resolution of GE imagery allows clear visualization of S. alterniflora texture and structure, facilitating the accurate delineation of boundaries, internal structure, and small patches. Second, GE imagery provides a more robust foundation for object-based classification. The object-based classification method used in this study relies on OBIA for effective image segmentation, and the quality of segmentation directly affects classification accuracy (Hao et al., 2021). Compared with Sentinel-2, GE imagery more accurately delineates object boundaries and internal structure, enabling the construction of clearly defined and spatially consistent segments that provide a superior basis for classification.
Moreover, the object-based classification applied in this study outperformed the pixel-based method in both accuracy and boundary delineation (Fig. 7). On the one hand, object-based classification reduces uncertainty in identifying S. alterniflora. The GE imagery contains complex spectral and texture information, which increases salt-and-pepper noise and reduces the reliability of pixel-level classification. By aggregating spectrally, texturally, and spatially consistent pixels, object-based classification reduces intra-class noise, improving both robustness and accuracy. On the other hand, the multi-scale optimized segmentation enhanced boundary delineation. The coexistence of S. alterniflora with native vegetation such as mangroves and Phragmites australis causes spectral mixing at patch edges, which pixel-based methods struggle to resolve. Although object-based methods improve boundary delineation, segmentation quality remains critical to classification accuracy. Therefore, we proposed a multi-scale optimized segmentation strategy designed to address the spectral heterogeneity at patch boundaries and spectral homogeneity within interiors, aiming to enhance the precision of boundary extraction. Specifically, a coarse scale was applied to homogeneous interiors, while a finer scale was used for complex boundaries. This strategy improves boundary delineation and avoids over-segmentation in uniform areas, enhancing both classification efficiency and accuracy.
4.2 The advantages of CM-SSM
This study produced the first Sub-meter S. alterniflora Map for Mainland China in 2020 (CM-SSM). Compared with the latest 10 m product (CMSA), CM-SSM demonstrated superior performance in classification accuracy, spatial detail representation, and spatial distribution statistics. Specifically, CM-SSM achieved higher classification accuracy, with OA and F1-score improved by 14.60 % and 0.24, respectively (Table 5). These advancements are primarily attributed to the development of sub-meter phenological features for S. alterniflora and the adoption of an object-based classification strategy (see Sect. 4.1).
CM-SSM also exhibited significant advantages in spatial detail extraction. Previous studies have indicated that S. alterniflora tends to exhibit a fragmented distribution pattern, where numerous small patches are often overlooked due to the limitations of 10 m resolution imagery (Zhou et al., 2024). Additionally, the species is often associated with diverse companion species and interspersed with narrow tidal creeks, which are difficult to distinguish using 10 m resolution due to the mixed pixel problem (Li et al., 2024). By increasing the mapping resolution to 0.9 m, CM-SSM effectively alleviated the impact of mixed pixels, enabling the accurate identification of small S. alterniflora patches, as well as their boundaries and internal structures (Fig. 8). This capability provides critical support for developing invasive species management strategies. For example, areas with dense small patches typically present higher risks of spread, and prioritizing these regions for control measures may help suppress expansion trends (Liu et al., 2016). Furthermore, CM-SSM's ability to delineate growth boundaries between S. alterniflora and companion species offers valuable insights into its invasion processes and potential threats to native species, such as mangroves (Chen and Shi, 2023). In addition, the accurate recognition of tidal creeks within S. alterniflora communities provides a data foundation for analyzing the relationship between creek morphology and spatial expansion (Kearney and Fagherazzi, 2016; Sanderson et al., 2000).
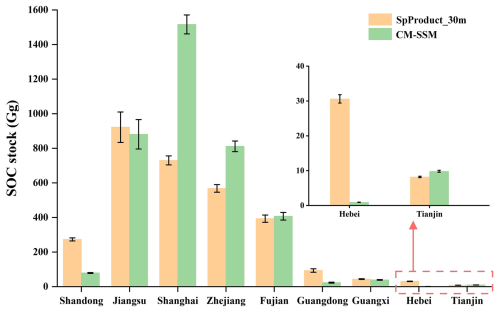
Figure 14Estimated SOC stock in the 0–1 m soil layer of S. alterniflora across coastal provinces of mainland China in 2020 based on SpProduct_30m and CM-SSM.
In terms of spatial distribution statistics, CM-SSM demonstrated higher accuracy. As shown in Sect. 3.2.3, although CM-SSM identified a significantly greater number of small S. alterniflora patches than CMSA, both products exhibited high consistency in detecting large S. alterniflora communities, resulting in a relatively small difference of only 1365 ha in total area (Fig. 9). However, spatial overlay analysis revealed a substantial spatial discrepancy of up to 34 273 ha between the two products, accounting for 57.73 % of the total CM-SSM area (Figs. 10 and 11). This discrepancy is primarily due to the limited spatial resolution of CMSA, which introduces classification errors. On the one hand, mixed pixels result in misclassification of other land types (e.g., creeks and open spaces) as S. alterniflora, leading to area overestimation. On the other hand, the failure to detect highly fragmented small patches leads to area underestimation. These findings underscore the critical role of spatial resolution in accurately capturing the distribution of S. alterniflora and highlight the limitations of relying solely on total area statistics, which may obscure substantial differences between products.
Accurate spatial distribution data of S. alterniflora are essential for the reliable quantification of its carbon storage (Xia et al., 2021; Xu et al., 2022). In this study, Soil Organic Carbon (SOC) storage was estimated using both CM-SSM and a multi-temporal 30 m dataset developed by Mao et al. (2019), based on the unified provincial-level SOC unit storage coefficient for S. alterniflora in 2020, as established by Zhang et al. (2024) through the integration of field measurements and spatial interpolation (Table S6). The results indicated that the total SOC storage estimated from CM-SSM reached 3767.69 Gg, which is 706.69 Gg higher than the estimate derived from the 30 m product, with particularly notable differences observed in Shanghai, Zhejiang, and Shandong (Fig. 14). The primary reason for this discrepancy lies in the resolution limitations of the 30 m product, which resulted in omission of small patches and misclassification at the edges, thereby reducing mapping accuracy. In contrast, CM-SSM provided a more accurate representation of the invasion pattern of S. alterniflora, leading to more reliable SOC estimates. This result highlights the value of fine-scale mapping in improving the accuracy of SOC storage estimation.
4.3 Limitations and prospects
Although this study proposed the OSPPF method and successfully produced the first Sub-meter S. alterniflora Map in Mainland China for the year 2020 (CM-SSM), several limitations remain. The OSPPF method has three main limitations. First, the automation of sample points generation requires further improvement. In this study, sample points were generated through visual interpretation based on field surveys and VHR imagery. However, this process is highly dependent on the expertise of researchers and is time-consuming, particularly for large-scale mapping. Although the ASC-CKM method proposed by Tian et al. (2025) leveraged the CascadeKMeans algorithm and the Mangrove Vegetation Index (MVI) for automated sample generation and achieved success in sub-meter mangrove mapping across China, an efficient and widely accepted classification index specific to S. alterniflora is currently lacking, which limits the direct transferability of this approach. Future research could focus on developing a highly discriminative spectral index tailored for S. alterniflora, with potential to support automated sample generation.
Additionally, the reliance on high-quality GE imagery constrains the broader applicability of the method. The frequent cloud cover of intertidal zones poses significant challenges to acquiring high-quality GE imagery at global scales or over long time series for sub-meter mapping of S. alterniflora. To address this issue, super-resolution techniques, which reconstruct high-resolution details from low-resolution imagery, have shown promising potential (Chen et al., 2024).
Third, the object-based classification incorporating multi-scale optimized segmentation still relies on existing large-scale S. alterniflora products during implementation. When applied to global-scale or long-term mapping, the reliance on global reference products presents a key limitation. To improve the transferability of the OSPPF method in regions lacking prior S. alterniflora products, alternative approaches can be adopted, such as generating an initial S. alterniflora mask using vegetation indices (e.g., NDVI, EVI), or automatically identifying potential distribution areas based on tidal zones, topography, and other environmental variables. In addition, future research may explore the application of deep learning models to S. alterniflora mapping, potentially replacing the object-based approach (Li et al., 2024). It is worth noting that the CM-SSM product developed in this study has the potential to serve as a valuable benchmark dataset for the training and evaluation of deep learning models.
The CM-SSM product has two limitations. First, temporal inconsistency among imagery sources may lead to classification errors. Although a comparison with imagery from adjacent years confirmed that the spatiotemporal dynamics of S. alterniflora were minor in most areas, and regions with temporal discrepancies accounted for only 2.20 % of the total area, temporal inconsistencies in imagery may still introduce slight errors. To address this issue, the integration of multi-source remote sensing data could be explored in future studies to mitigate the impact of temporal mismatches and further reduce classification errors. Second, manual visual interpretation introduces subjectivity, which may result in minor inaccuracies. Although this study utilized VHR imagery for visual interpretation to optimize the mapping product, differences in S. alterniflora texture, morphology, and spectral appearance across climate zones posed challenges for consistent manual interpretation. Future research could explore semi-automated methods or interpretation strategies supported by deep learning to reduce uncertainties arising from human intervention.
The 2020 Sub-meter S. alterniflora Map of Mainland China (CM-SSM) generated by this study is openly available at https://doi.org/10.5281/zenodo.16296823 (Xu et al., 2025).
This study proposed a novel Object- and Sub-meter-enhanced Pixel-based Phenological Feature (OSPPF) composite method to generate the first 2020 Sub-meter S. alterniflora Map of Mainland China (CM-SSM). The OSPPF method integrates multi-source remote sensing imagery and employs an object-based classification method with a multi-scale optimized segmentation strategy, effectively addressing limitations of existing methods in delineating small patches, boundaries, and internal details of S. alterniflora. Compared with the latest 10 m resolution S. alterniflora map (CMSA), the CM-SSM shows significant improvements in classification accuracy, spatial detail, and spatial distribution statistics. Specifically, OA and F1 score of CM-SSM are 14.60 % and 0.24 higher than those of CMSA, respectively. While the total area difference between the two products is only 2.30 %, spatial distribution discrepancies reach 57.73 %, and the number of detected patches in CM-SSM is 17 times greater than in CMSA. The CM-SSM product and its underlying OSPPF method provide high-precision baseline data for monitoring S. alterniflora, and offer a scalable framework for future sub-meter mapping at broader spatial and temporal scales. These advancements hold substantial potential for supporting S. alterniflora management effectiveness assessments and blue carbon stock estimations.
The supplement related to this article is available online at https://doi.org/10.5194/essd-17-6601-2025-supplement.
BZ, MX, and JT designed the research and wrote the manuscript. MJ, DM, and KC supervised and reviewed the paper. XZ, HJ, JS, YH, and MW collected and processed the data. YK, ZZ, LZ, XL, and HG reviewed the manuscript. BZ and MX contributed equally to this work.
The contact author has declared that none of the authors has any competing interests.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. While Copernicus Publications makes every effort to include appropriate place names, the final responsibility lies with the authors. Views expressed in the text are those of the authors and do not necessarily reflect the views of the publisher.
We thank the editor and the reviewers for their valuable comments.
This research has been supported by the National Natural Science Foundation of China (grant nos. 42171330 and 42571440) and the Natural Science Foundation of Beijing Municipality (grant no. L251047).
This paper was edited by Yuanzhi Yao and reviewed by two anonymous referees.
Ai, J., Gao, W., Gao, Z., Shi, R., and Zhang, C.: Phenology-based Spartina alterniflora mapping in coastal wetland of the Yangtze Estuary using time series of GaoFen satellite no. 1 wide field of view imagery, J. Appl. Remote Sens., 11, 026020–026020, https://doi.org/10.1117/1.JRS.11.026020, 2017.
Breiman, L.: Random forests, Mach. Learn., 45, 5–32, https://doi.org/10.1023/A:1010933404324, 2001.
Chandrasekar, K., Sesha Sai, M., Roy, P., and Dwevedi, R.: Land Surface Water Index (LSWI) response to rainfall and NDVI using the MODIS Vegetation Index product, Int. J. Remote Sens., 31, 3987–4005, https://doi.org/10.1080/01431160802575653, 2010.
Chen, M., Ke, Y., Bai, J., Li, P., Lyu, M., Gong, Z., and Zhou, D.: Monitoring early stage invasion of exotic Spartina alterniflora using deep-learning super-resolution techniques based on multisource high-resolution satellite imagery: A case study in the Yellow River Delta, China, Int. J. Appl. Earth Obs. Geoinf., 92, 102180, https://doi.org/10.1016/j.jag.2020.102180, 2020.
Chen, W. and Shi, C.: Fine-scale mapping of Spartina alterniflora-invaded mangrove forests with multi-temporal WorldView-Sentinel-2 data fusion, Remote Sens. Environ., 295, 113690, https://doi.org/10.1016/j.rse.2023.113690, 2023.
Chen, W., Tian, J., Song, J., Li, X., Ke, Y., Zhu, L., Yu, Y., Ou, Y., and Gong, H.: A noval super-resolution model for 10-m mangrove mapping with landsat-5, IEEE Trans. Geosci. Remote Sens., https://doi.org/10.1109/TGRS.2024.3407363, 2024.
Chen, Y. and Kirwan, M. L.: A phenology-and trend-based approach for accurate mapping of sea-level driven coastal forest retreat, Remote Sens. Environ., 281, 113229, https://doi.org/10.1016/j.rse.2022.113229, 2022.
Chen, Y., Tian, J., Song, J., Chen, W., Zhou, B., Qu, X., and Zhang, L.: Developing a new index with time series Sentinel-2 for accurate tidal flats mapping in China, Sci. Total Environ., 958, 178037, https://doi.org/10.1016/j.scitotenv.2024.178037, 2025.
Dong, D., Huang, H., and Gao, Q.: Tracking the Dynamics of Spartina alterniflora with WorldView-2/3 and Sentinel-1/2 Imagery in Zhangjiang Estuary, China, Water, 16, 1780, https://doi.org/10.3390/w16131780, 2024.
Drăguţ, L., Tiede, D., and Levick, S. R.: ESP: a tool to estimate scale parameter for multiresolution image segmentation of remotely sensed data, Int. J. Geogr. Inf. Sci., 24, 859–871, https://doi.org/10.1080/13658810903174803, 2010.
Dronova, I.: Object-based image analysis in wetland research: A review, Remote Sens., 7, 6380–6413, https://doi.org/10.3390/rs70506380, 2015.
Du, M. and Noguchi, N.: Monitoring of wheat growth status and mapping of wheat yield's within-field spatial variations using color images acquired from UAV-camera system, Remote Sens., 9, 289, https://doi.org/10.3390/rs9030289, 2017.
Frohn, R., Autrey, B., Lane, C., and Reif, M.: Segmentation and object-oriented classification of wetlands in a karst Florida landscape using multi-season Landsat-7 ETM+ imagery, Int. J. Remote Sens., 32, 1471–1489, https://doi.org/10.1080/01431160903559762, 2011.
Gao, B.-C.: NDWI – A normalized difference water index for remote sensing of vegetation liquid water from space, Remote Sens. Environ., 58, 257–266, https://doi.org/10.1016/S0034-4257(96)00067-3, 1996.
Gitelson, A. A., Kaufman, Y. J., Stark, R., and Rundquist, D.: Novel algorithms for remote estimation of vegetation fraction, Remote Sens. Environ., 80, 76–87, https://doi.org/10.1016/S0034-4257(01)00289-9, 2002.
Hao, S., Cui, Y., and Wang, J.: Segmentation scale effect analysis in the object-oriented method of high-spatial-resolution image classification, Sensors, 21, 7935, https://doi.org/10.3390/s21237935, 2021.
Haralick, R. M., Shanmugam, K., and Dinstein, I. H.: Textural features for image classification, IEEE T. Syst. Man Cybernet., 610–621, https://doi.org/10.1109/TSMC.1973.4309314, 2007.
Hu, Y., Tian, B., Yuan, L., Li, X., Huang, Y., Shi, R., Jiang, X., and Sun, C.: Mapping coastal salt marshes in China using time series of Sentinel-1 SAR, ISPRS J. Photogramm. Remote Sens., 173, 122–134, https://doi.org/10.1016/j.isprsjprs.2021.01.003, 2021.
Huete, A., Didan, K., Miura, T., Rodriguez, E. P., Gao, X., and Ferreira, L. G.: Overview of the radiometric and biophysical performance of the MODIS vegetation indices, Remote Sens. Environ., 83, 195–213, https://doi.org/10.1016/S0034-4257(02)00096-2, 2002.
Jackson, M. V., Fuller, R. A., Gan, X., Li, J., Mao, D., Melville, D. S., Murray, N. J., Wang, Z., and Choi, C.-Y.: Dual threat of tidal flat loss and invasive Spartina alterniflora endanger important shorebird habitat in coastal mainland China, J. Environ. Manage., 278, 111549, https://doi.org/10.1016/j.jenvman.2020.111549, 2021.
Kearney, W. S. and Fagherazzi, S.: Salt marsh vegetation promotes efficient tidal channel networks, Nat. Commun., 7, 12287, https://doi.org/10.1038/ncomms12287, 2016.
Li, B., Liao, C.-H., Zhang, X.-D., Chen, H.-l., Wang, Q., Chen, Z.-Y., Gan, X.-J., Wu, J.-H., Zhao, B., Ma, Z.-J., Cheng, X.-l., Jiang, L.-F., and Chen, J.-K.: Spartina alterniflora invasions in the Yangtze River estuary, China: an overview of current status and ecosystem effects, Ecol. Eng., 35, 511–520, https://doi.org/10.1016/j.ecoleng.2008.05.013, 2009.
Li, H., Han, Y., and Chen, J.: Combination of Google Earth imagery and Sentinel-2 data for mangrove species mapping, J. Appl. Remote Sens., 14, 010501, https://doi.org/10.1117/1.JRS.14.010501, 2020.
Li, H., Mao, D., Wang, Z., Huang, X., Li, L., and Jia, M.: Invasion of Spartina alterniflora in the coastal zone of mainland China: Control achievements from 2015 to 2020 towards the Sustainable Development Goals, J. Environ. Manage., 323, 116242, https://doi.org/10.1016/j.jenvman.2022.116242, 2022a.
Li, L., Liu, W., Tao, Y., Xu, X., Fu, W., and Dong, J.: Diffusion dynamics and driving forces of Spartina alterniflora in the Guangxi Shankou Mangrove Reserve, Acta Ecol. Sin., 41, 6814–6824, https://doi.org/10.5846/stxb202101260275, 2021.
Li, X., Tian, J., Li, X., Wang, L., Gong, H., Shi, C., Nie, S., Zhu, L., Chen, B., and Pan, Y.: Developing a sub-meter phenological spectral feature for mapping poplars and willows in urban environment, ISPRS J. Photogramm. Remote Sens., 193, 77–89, https://doi.org/10.1016/j.isprsjprs.2022.09.002, 2022b.
Li, X., Tian, J., Li, X., Yu, Y., Ou, Y., Zhu, L., Zhu, X., Zhou, B., and Gong, H.: Annual mapping of Spartina alterniflora with deep learning and spectral-phenological features from 2017 to 2021 in the mainland of China, Int. J. Remote Sens., 45, 3172–3199, https://doi.org/10.1080/01431161.2024.2343136, 2024.
Liu, C., Jiang, H., Zhang, S., Li, C., Pan, X., Lu, J., and Hou, Y.: Expansion and management implications of invasive alien Spartina alterniflora in Yancheng salt marshes, China, Open J. Ecol., 6, 113–128, https://doi.org/10.4236/oje.2016.63012, 2016.
Liu, M., Li, H., Li, L., Man, W., Jia, M., Wang, Z., and Lu, C.: Monitoring the invasion of Spartina alterniflora using multi-source high-resolution imagery in the Zhangjiang Estuary, China, Remote Sens., 9, 539, https://doi.org/10.3390/rs9060539, 2017a.
Liu, M., Mao, D., Wang, Z., Li, L., Man, W., Jia, M., Ren, C., and Zhang, Y.: Rapid invasion of Spartina alterniflora in the coastal zone of mainland China: new observations from Landsat OLI images, Remote Sens., 10, 1933, https://doi.org/10.3390/rs10121933, 2018.
Liu, W., Chen, X., Strong, D. R., Pennings, S. C., Kirwan, M. L., Chen, X., and Zhang, Y.: Climate and geographic adaptation drive latitudinal clines in biomass of a widespread saltmarsh plant in its native and introduced ranges, Limnol. Oceanogr., 65, 1399–1409, https://doi.org/10.1002/lno.11395, 2020.
Liu, X., Liu, H., Gong, H., Lin, Z., and Lv, S.: Appling the one-class classification method of maxent to detect an invasive plant Spartina alterniflora with time-series analysis, Remote Sens., 9, 1120, https://doi.org/10.3390/rs9111120, 2017b.
Lourenço, P., Teodoro, A. C., Gonçalves, J. A., Honrado, J. P., Cunha, M., and Sillero, N.: Assessing the performance of different OBIA software approaches for mapping invasive alien plants along roads with remote sensing data, Int. J. Appl. Earth Obs. Geoinf., 95, 102263, https://doi.org/10.1016/j.jag.2020.102263, 2021.
Lv, J., Jiang, W., Wang, W., Wu, Z., Liu, Y., Wang, X., and Li, Z.: Wetland loss identification and evaluation based on landscape and remote sensing indices in Xiong'an new area, Remote Sens., 11, 2834, https://doi.org/10.3390/rs11232834, 2019.
Mancino, G., Ferrara, A., Padula, A., and Nolè, A.: Cross-comparison between Landsat 8 (OLI) and Landsat 7 (ETM+) derived vegetation indices in a Mediterranean environment, Remote Sens., 12, 291, https://doi.org/10.3390/rs12020291, 2020.
Mao, D., Liu, M., Wang, Z., Li, L., Man, W., Jia, M., and Zhang, Y.: Rapid invasion of Spartina alterniflora in the coastal zone of mainland China: Spatiotemporal patterns and human prevention, Sensors, 19, 2308, https://doi.org/10.3390/s19102308, 2019.
McNemar, Q.: Note on the sampling error of the difference between correlated proportions or percentages, Psychometrika, 12, 153–157, https://doi.org/10.1007/BF02295996, 1947.
Meng, W., Feagin, R. A., Innocenti, R. A., Hu, B., He, M., and Li, H.: Invasion and ecological effects of exotic smooth cordgrass Spartina alterniflora in China, Ecol. Eng., 143, 105670, https://doi.org/10.1016/j.ecoleng.2019.105670, 2020.
Merzlyak, M. N., Gitelson, A. A., Chivkunova, O. B., and Rakitin, V. Y.: Non-optical detection of pigment changes during leaf senescence and fruit ripening, Physiol. Plant., 106, 135–141, https://doi.org/10.1034/j.1399-3054.1999.106119.x, 1999.
Ni, R., Tian, J., Li, X., Yin, D., Li, J., Gong, H., Zhang, J., Zhu, L., and Wu, D.: An enhanced pixel-based phenological feature for accurate paddy rice mapping with Sentinel-2 imagery in Google Earth Engine, ISPRS J. Photogramm. Remote Sens., 178, 282–296, https://doi.org/10.1016/j.isprsjprs.2021.06.018, 2021.
Okoye, O. K., Li, H., and Gong, Z.: Retraction of invasive Spartina alterniflora and its effect on the habitat loss of endangered migratory bird species and their decline in YNNR using remote sensing technology, Ecol. Evol., 10, 13810–13824, https://doi.org/10.1002/ece3.6971, 2020.
Ouyang, Z.-T., Gao, Y., Xie, X., Guo, H.-Q., Zhang, T.-T., and Zhao, B.: Spectral discrimination of the invasive plant Spartina alterniflora at multiple phenological stages in a saltmarsh wetland, PLoS One, 8, e67315, https://doi.org/10.1371/journal.pone.0067315, 2013.
Rouse, J., Haas, R., Schell, J., and Deering, D.: Monitoring vegetation systems in the Great Plains with ERTS. 1974, in: Proceeding Of Erts-Symposium, https://ntrs.nasa.gov/citations/19740022614 (last access: 25 November 2025), 1974.
Sanderson, E. W., Ustin, S. L., and Foin, T. C.: The influence of tidal channels on the distribution of salt marsh plant species in Petaluma Marsh, CA, USA, Plant Ecol., 146, 29–41, https://doi.org/10.1023/A:1009882110988, 2000.
Savitzky, A. and Golay, M. J.: Smoothing and differentiation of data by simplified least squares procedures, Anal. Chem., 36, 1627–1639, https://doi.org/10.1021/ac60214a047, 1964.
Sun, C., Liu, Y., Zhao, S., Zhou, M., Yang, Y., and Li, F.: Classification mapping and species identification of salt marshes based on a short-time interval NDVI time-series from HJ-1 optical imagery, Int. J. Appl. Earth Obs. Geoinf., 45, 27–41, https://doi.org/10.1016/j.jag.2015.10.008, 2016.
Sun, C., Li, J., Liu, Y., Liu, Y., and Liu, R.: Plant species classification in salt marshes using phenological parameters derived from Sentinel-2 pixel-differential time-series, Remote Sens. Environ., 256, 112320, https://doi.org/10.1016/j.rse.2021.112320, 2021.
Sun, L., Shao, D., Xie, T., Gao, W., Ma, X., Ning, Z., and Cui, B.: How does spartina alterniflora invade in salt marsh in relation to Tidal Channel networks? Patterns and processes, Remote Sens., 12, 2983, https://doi.org/10.3390/rs12182983, 2020.
Tian, J., Wang, L., Yin, D., Li, X., Diao, C., Gong, H., Shi, C., Menenti, M., Ge, Y., Nie, S., Ou, Y., Song, X., and Liu, X.: Development of spectral-phenological features for deep learning to understand Spartina alterniflora invasion, Remote Sens. Environ., 242, 111745, https://doi.org/10.1016/j.rse.2020.111745, 2020a.
Tian, J., Wang, L., Diao, C., Zhang, Y., Jia, M., Zhu, L., Xu, M., Li, X., and Gong, H.: National scale sub-meter mangrove mapping using an augmented border training sample method, ISPRS J. Photogramm. Remote Sens., 220, 156–171, https://doi.org/10.1016/j.isprsjprs.2024.12.009, 2025.
Tian, Y., Jia, M., Wang, Z., Mao, D., Du, B., and Wang, C.: Monitoring invasion process of Spartina alterniflora by seasonal Sentinel-2 imagery and an object-based random forest classification, Remote Sens., 12, 1383, https://doi.org/10.3390/rs12091383, 2020b.
Wan, H., Wang, Q., Jiang, D., Fu, J., Yang, Y., and Liu, X.: Monitoring the invasion of Spartina alterniflora using very high resolution unmanned aerial vehicle imagery in Beihai, Guangxi (China), Scient. World J., 2014, 638296, https://doi.org/10.1155/2014/638296, 2014.
Wang, A., Chen, J., Jing, C., Ye, G., Wu, J., Huang, Z., and Zhou, C.: Monitoring the invasion of Spartina alterniflora from 1993 to 2014 with Landsat TM and SPOT 6 satellite data in Yueqing Bay, China, PLoS One, 10, e0135538, https://doi.org/10.1371/journal.pone.0135538, 2015a.
Wang, T., Zhang, H., Lin, H., and Fang, C.: Textural–spectral feature-based species classification of mangroves in Mai Po Nature Reserve from Worldview-3 imagery, Remote Sens., 8, 24, https://doi.org/10.3390/rs8010024, 2015b.
Wang, X., Xiao, X., Zou, Z., Hou, L., Qin, Y., Dong, J., Doughty, R. B., Chen, B., Zhang, X., Chen, Y., Ma, J., Zhao, B., and Li, B.: Mapping coastal wetlands of China using time series Landsat images in 2018 and Google Earth Engine, ISPRS J. Photogramm. Remote Sens., 163, 312–326, https://doi.org/10.1016/j.isprsjprs.2020.03.014, 2020.
Wang, X., Wang, L., Tian, J., and Shi, C.: Object-based spectral-phenological features for mapping invasive Spartina alterniflora, Int. J. Appl. Earth Obs. Geoinf., 101, 102349, https://doi.org/10.1016/j.jag.2021.102349, 2021.
Windle, A. E., Staver, L. W., Elmore, A. J., Scherer, S., Keller, S., Malmgren, B., and Silsbe, G. M.: Multi-temporal high-resolution marsh vegetation mapping using unoccupied aircraft system remote sensing and machine learning, Front. Remote Sens., 4, 1140999, https://doi.org/10.3389/frsen.2023.1140999, 2023.
Wu, Y., Xiao, X., Chen, B., Ma, J., Wang, X., Zhang, Y., Zhao, B., and Li, B.: Tracking the phenology and expansion of Spartina alterniflora coastal wetland by time series MODIS and Landsat images, Multimed. Tools Appl., 79, 5175–5195, https://doi.org/10.1007/s11042-018-6314-9, 2020.
Xia, S., Wang, W., Song, Z., Kuzyakov, Y., Guo, L., Van Zwieten, L., Li, Q., Hartley, I. P., Yang, Y., Wang, Y., Quine, T. A., Liu, C., and Wang, H.: Spartina alterniflora invasion controls organic carbon stocks in coastal marsh and mangrove soils across tropics and subtropics, Global Change Biol., 27, 1627–1644, https://doi.org/10.1111/gcb.15516, 2021.
Xu, M., Tian, J., and Zhou, B.: National-Scale Sub-meter Mapping of Spartina alterniflora in Mainland China 2020, Zenodo [data set], https://doi.org/10.5281/zenodo.16296823, 2025.
Xu, X., Wei, S., Chen, H., Li, B., and Nie, M.: Effects of Spartina invasion on the soil organic carbon content in salt marsh and mangrove ecosystems in China, J. Appl. Ecol., 59, 1937–1946, https://doi.org/10.1111/1365-2664.14202, 2022.
Yan, D., Li, J., Yao, X., and Luan, Z.: Quantifying the long-term expansion and dieback of Spartina alterniflora using google earth engine and object-based hierarchical random forest classification, IEEE J. Select. Top. Appl. Earth Obs. Remote Sens., 14, 9781–9793, https://doi.org/10.1109/JSTARS.2021.3114116, 2021.
Zeng, J., Sun, Y., Cao, P., and Wang, H.: A phenology-based vegetation index classification (PVC) algorithm for coastal salt marshes using Landsat 8 images, Int. J. Appl. Earth Obs. Geoinf., 110, 102776, https://doi.org/10.1016/j.jag.2022.102776, 2022.
Zhang, D., Hu, Y., Liu, M., Chang, Y., Yan, X., Bu, R., Zhao, D., and Li, Z.: Introduction and spread of an exotic plant, Spartina alterniflora, along coastal marshes of China, Wetlands, 37, 1181–1193, https://doi.org/10.1007/s13157-017-0950-0, 2017.
Zhang, J., Mao, D., Liu, J., Chen, Y., Kirwan, M., Sanders, C., Zhou, J., Lu, Z., Qin, G., Huang, X., Li, H., Yan, H., Jiao, N., Su, J., and Wang, F.: Spartina alterniflora invasion benefits blue carbon sequestration in China, Sci. Bull., 69, 1991–2000, https://doi.org/10.1016/j.scib.2024.04.049, 2024.
Zhang, X., Xiao, X., Wang, X., Xu, X., Chen, B., Wang, J., Ma, J., Zhao, B., and Li, B.: Quantifying expansion and removal of Spartina alterniflora on Chongming island, China, using time series Landsat images during 1995–2018, Remote Sens. Environ., 247, 111916, https://doi.org/10.1016/j.rse.2020.111916, 2020.
Zhang, X., Xiao, X., Qiu, S., Xu, X., Wang, X., Chang, Q., Wu, J., and Li, B.: Quantifying latitudinal variation in land surface phenology of Spartina alterniflora saltmarshes across coastal wetlands in China by Landsat 7/8 and Sentinel-2 images, Remote Sens. Environ., 269, 112810, https://doi.org/10.1016/j.rse.2021.112810, 2022.
Zhang, X., Liu, L., Zhao, T., Chen, X., Lin, S., Wang, J., Mi, J., and Liu, W.: GWL_FCS30: a global 30 m wetland map with a fine classification system using multi-sourced and time-series remote sensing imagery in 2020, Earth Syst. Sci. Data, 15, 265–293, https://doi.org/10.5194/essd-15-265-2023, 2023a.
Zhang, X., Xiao, X., Wang, X., Xu, X., Qiu, S., Pan, L., Ma, J., Ju, R., Wu, J., and Li, B.: Continual expansion of Spartina alterniflora in the temperate and subtropical coastal zones of China during 1985–2020, Int. J. Appl. Earth Obs. Geoinf., 117, 103192, https://doi.org/10.1016/j.jag.2023.103192, 2023b.
Zhao, C., Jia, M., Wang, Z., Mao, D., and Wang, Y.: Toward a better understanding of coastal salt marsh mapping: A case from China using dual-temporal images, Remote Sens. Environ., 295, 113664, https://doi.org/10.1016/j.rse.2023.113664, 2023.
Zhao, Z., Yuan, L., Li, W., Tian, B., and Zhang, L.: Re-invasion of Spartina alterniflora in restored saltmarshes: Seed arrival, retention, germination, and establishment, J. Environ. Manage., 266, 110631, https://doi.org/10.1016/j.jenvman.2020.110631, 2020.
Zheng, J., Wei, H., Chen, R., Liu, J., Wang, L., and Gu, W.: Invasive trends of Spartina Alterniflora in the southeastern Coast of China and potential distributional impacts on mangrove forests, Plants, 12, 1923, https://doi.org/10.3390/plants12101923, 2023b.
Zheng, J.-Y., Hao, Y.-Y., Wang, Y.-C., Zhou, S.-Q., Wu, W.-B., Yuan, Q., Gao, Y., Guo, H.-Q., Cai, X.-X., and Zhao, B.: Coastal wetland vegetation classification using pixel-based, object-based and deep learning methods based on RGB-UAV, Land, 11, 2039, https://doi.org/10.3390/land11112039, 2022.
Zheng, X., Javed, Z., Liu, B., Zhong, S., Cheng, Z., Rehman, A., Du, D., and Li, J.: Impact of Spartina alterniflora invasion in coastal wetlands of China: boon or bane?, Biology, 12, 1057, https://doi.org/10.3390/biology12081057, 2023a.
Zhou, B., Xu, M., Tian, J., Huang, Y., Song, J., Zhu, L., Zhu, X., Qu, X., Zhang, L., Li, X., and Gong, H.: Mapping the invasive Spartina alterniflora in sub-meter level with improved phenological spectral features and deep learning method, Int. J. Digit. Earth, 17, 2434634, https://doi.org/10.1080/17538947.2024.2434634, 2024.
Zuo, P., Zhao, S., Liu, C. a., Wang, C., and Liang, Y.: Distribution of Spartina spp. along China's coast, Ecol. Eng., 40, 160–166, https://doi.org/10.1016/j.ecoleng.2011.12.014, 2012.
Zuo, Y., Yang, G., Sun, W., Huang, K., Yang, S., Chen, B., Wang, L., Meng, X., Wang, Y., Li, J., and Zhan, Y.: SAI: A Spartina alterniflora index based on Sentinel-2 multispectral imagery for Spartina alterniflora mapping, J. Remote Sens., https://doi.org/10.34133/remotesensing.0510, 2025.