the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Toward better conservation: a spatial analysis of species occurrence data from the Global Biodiversity Information Facility
Susmita Dasgupta
Brian Blankespoor
David Wheeler
The world is facing an unprecedented loss of biodiversity, with nearly one million species on the brink of extinction, and the extinction rate accelerating. Conservation efforts are often hindered by insufficient information on crucial ecosystems. To address this issue, our paper leverages advances in machine-based pattern recognition to estimate species occurrence maps using georeferenced data from the Global Biodiversity Information Facility (GBIF). Our algorithms have generated maps for more than 600 000 species, including vertebrates, arthropods, mollusks, other animals, vascular plants, fungi, and other organisms. Validation involved comparing these maps with expert maps for mammals, ants, and vascular plants. We found a close similarity in global distribution patterns, with regional differences attributed to technical variations or necessary revisions in existing expert maps based on GBIF data. As a practical application, we identified the global distributions of approximately 68 000 species with small ranges (25 km × 25 km or less) confined to a single country. Our maps reveal a skewed international distribution of these species, identifying 30 countries where 78.2 % are concentrated. These results highlight the need to integrate the newly mapped GBIF data into global conservation planning. Our algorithms support rapid updates and the creation of new maps as GBIF occurrence reports increase. The data are available on the World Bank Development Data Hub at https://doi.org/10.57966/h21e-vq42 (Dasgupta et al., 2024).
- Article
(8517 KB) - Full-text XML
- BibTeX
- EndNote
Toward better conservation: a spatial analysis of species occurrence data from the Global Biodiversity Information Facility © by International Bank for Reconstruction and Development/International Development Association or The World Bank.
This work is provided under a Creative Commons 4.0 Attribution International License, with the following mandatory and binding addition: Any and all disputes arising under this License that cannot be settled amicably shall be submitted to mediation in accordance with the WIPO Mediation Rules in effect at the time the work was published. If the request for mediation is not resolved within forty-five (45) days of the request, either You or the Licensor may, pursuant to a notice of arbitration communicated by reasonable means to the other party refer the dispute to final and binding arbitration to be conducted in accordance with UNCITRAL Arbitration Rules as then in force. The arbitral tribunal shall consist of a sole arbitrator and the language of the proceedings shall be English unless otherwise agreed. The place of arbitration shall be where the Licensor has its headquarters. The arbitral proceedings shall be conducted remotely (e.g., via telephone conference or written submissions) whenever practicable, or held at the World Bank headquarters in Washington DC.
The world is losing biodiversity at an unprecedented rate. One million plant and animal species may be near extinction, and the pace of extinction is accelerating. According to Pimm et al. (2014), the species extinction rate is at least one thousand times the background rate. The Living Planet Index (https://www.livingplanetindex.org/, last access: 26 January 2026), an indicator of global biodiversity based on population trends for vertebrate species in terrestrial, freshwater, and marine habitats, provides corroborating evidence, showing a 73 % decline since 1970 (WWF, 2024). In response to such alarming indicators, 188 governments in the Convention on Biological Diversity (CBD) ratified the Kunming-Montreal Global Biodiversity Framework (GBF) at the fifteenth meeting of the CBD's Conference of the Parties (COP 15) in December 2022. Among other measures, the GBF committed participants to protecting 30 % of global biodiversity by 2030 (UNEP, 2022). Effectively implementing the GBF requires an understanding of (1) the spatial distribution of global biodiversity to be protected and (2) how protecting 30 % of the planet can best conserve this biodiversity, taking the opportunity value of protected areas into account.
Unfortunately, conservation efforts worldwide are often hindered by limited information on critical ecosystems and biodiversity (Hughes et al., 2024). Comprehensive species coverage is significantly lacking; instead, the predominant focus is on vertebrates and vascular plants, neglecting crucial taxa like invertebrates and other major phyla. This gap creates a policy dilemma for meeting the 2030 GBF commitment of 30 % global protection. If global biodiversity assessments are limited to previously mapped species, policy makers and the conservation community will effectively ignore the enormous population of other species whose occurrences are reported by the Global Biodiversity Information Facility (GBIF). In short, policymakers cannot automatically assume that previously mapped species adequately represent the larger population, a point further discussed by Kass et al. (2022) in the case of invertebrates.
To help bridge the data gap, this study uses GBIF species occurrence records to revisit global biodiversity's spatial distribution. GBIF's reporting network has expanded over 15 years to include over 2 million species' occurrences, with a daily increase of about 1.3 million records in the last two years. Most of these records include locational coordinates, making it possible to estimate the spatial distribution of previously unmapped species, as well as improve estimates for those with existing maps. The algorithm that is presented and implemented in this study generates species maps directly from the GBIF occurrence data, updating the maps automatically as new occurrence data become available. Maps are generated for all species whose data satisfy our computational criteria (currently around 600 000 species). By overlaying the maps with a high-resolution grid, the view of global biodiversity is broadened from the traditional focus on vertebrate animals to encompass greater representation for invertebrates, other animals, plants, fungi, and other non-animal and non-plant species. The maps are then used to develop new indicators of species endemism and identify species with small, vulnerable habitats. Traditionally, species are considered endemic if they reside 100 % in one country; this study examines the effects of lowering this threshold to 95 % and 90 %. Additionally, since small-range status lacks a definitive minimum habitat size, the study explores different area sizes for species with limited occurrence regions.
The study's approach should be viewed as complementary to previous biodiversity assessments (e.g. Map of Life) and can benefit the policy process in several major ways. For existing mapped species, rapid updates using the study's algorithm can help to identify cases where newly reported occurrences suggest alteration of map boundaries. For unmapped species, the approach can provide new information useful for global biodiversity assessments. In addition, the mapping exercise can yield valuable insights on the global distribution of endemic species and small-range species that are especially vulnerable to human encroachment.
The open-access data, algorithm, and programming code developed in this study align with principles of open and equitable sharing of location-specific biodiversity information (Blankespoor et al., 2025; Tanalgo, 2025). It is hoped that this approach would facilitate coordination among countries and enable policymakers, researchers, and conservation practitioners to take informed, coordinated actions to safeguard biodiversity.
2.1 Data source and tools
Our data source is an international network funded by the world's governments that provides open access to data about all types of life on Earth. Other international organizations with which the GBIF collaborates include the Catalogue of Life (https://www.catalogueoflife.org/, last access: 26 January 2026) partnership, Biodiversity Information Standards, Consortium for the Barcode of Life (CBOL), Encyclopedia of Life (EOL), and Global Earth Observation System of Systems (GEOSS). The GBIF provides a continuously updated, open-source repository of geolocated, date-stamped reports of species occurrences from many institutions and nongovernmental organizations (NGOs) worldwide. These reports can be accessed directly from the GBIF's Occurrence and Maps application programming interfaces (APIs) at https://www.gbif.org/developer/occurrence (last access: 26 January 2026) and https://www.gbif.org/developer/maps (last access: 26 January 2026), respectively. Its georeferenced data for plants, animals, fungi, and microbes hold the potential for vastly expanding the species domain maps that provide a critical foundation for global conservation planning. Access to the GBIF's full database is offered through Google's BigQuery, Amazon Web Services (AWS), and other cloud-based services.
In this study, we use BigQuery to download the full GBIF database due to its convenient dataset size reduction tools (GBIF, 2024). We limit the GBIF occurrence data to geolocated reports since 1970 for species with at least five unique reporting locations, which our mapping algorithm requires. Since our mapping algorithm operates reliably on several thousand points at most, we cap the data at a maximum of 20 000 randomly selected reports per species to ensure reliability. This limit drastically reduces the size of the download dataset since some species have millions of occurrence records (e.g., the American robin [Turdus migratorius] currently has 21 258 907 reported occurrences). We accept the GBIF's protocols for occurrence report admissibility. Detailed descriptions of the GBIF's protocols and database elements can be found at https://www.gbif.org/data-quality-requirements-occurrences (last access: 26 January 2026).
2.2 Analysis
The premise of our analysis is that species maps underpin spatial analyses of global biodiversity. In theory, if all species are treated equally, a map of global biodiversity could be created by (1) choosing the best available map for each species; (2) overlaying all chosen maps on a high-resolution global grid; and (3) counting the total species incidence in each grid cell. In practice, however, global biodiversity analyses can modify species counting in several major ways. For example, species may be assigned weights, based on the branches they occupy in the “tree of life,” which describes overall genetic variation in the global biome. Also, species weights for conservation priority-setting may vary, based on the species' widely differing vulnerabilities to human encroachment. In addition, the distribution of a species may not be uniform in the spaces enclosed by its maps. If its spatial distribution density is known, its counting weight for each cell can be made proportional to its likelihood of occurrence in that cell. Abundant scientific literature offers examples of weighted species counting (e.g., Guo et al., 2022; Jenkins et al., 2015; Pimm et al., 2014; Veach et al., 2017); however, the requisite research may require detailed genetic and environmental data, as well as expert analysis of their roles in assigning counting weights. Inevitably, the intensive processes involved are time-consuming, requiring technical resources that are in short supply. As a result, a large gap has emerged between the population of species with GBIF occurrence records and that for which research-driven counting weights are available.
The mapping algorithm utilized in this study is a by-product of recent advances in machine-based pattern recognition, cluster analysis, and image processing. In terms of computational geometry, it addresses the problem of efficient bounding of a spatial set, given a subset of actually observed points. Traditional algorithms that draw simple convex hulls poorly represent sets with irregular shapes as a polygon is considered convex if none of its corners bend inward and a convex hull is the smallest convex polygon that encloses all points in a set. In contrast, this study's alphahull algorithm, developed by Pateiro-López and Rodríguez-Casal (2010), which is a function in the R programming language can construct continuous non-convex boundaries for efficient representation. This powerful feature has motivated alphahull's rapid adoption for species range analysis (Guo et al., 2022; Kass et al., 2022).
In our study, alphahull successfully estimates occurrence maps for 92.9 % (567 464) of the 610 694 species in our database. For each of the remaining 7.1 % of species in the database, we employ a standard k-means algorithm to separate occurrence reports into spatial clusters and draw a convex hull around each. We assess the utility of these mapping algorithms for delineating species occurrence regions, with illustrative examples provided in the Annex. Our algorithms estimate occurrence maps for terrestrial, coastal, and marine species.
2.3 Spatial selection bias
We acknowledge that GBIF occurrence reports are often produced by voluntary exercises that do not utilize scientific sampling methods. As a result, spatial point densities in species occurrence reports are positively related to physical accessibility, population density, and income (Borgelt et al., 2022; Garcia-Roselló et al., 2023; Isaac and Pocock, 2015; Reddy and Dávalos, 2003). This means that (all else being equal) species sightings are more likely to occur in areas (1) near transport arteries; (2) with a greater number of inhabitants to identify species, and (3) where more inhabitants have enough disposable income to support species search and reporting costs.
These factors complicate attempts to map species population densities from occurrence reports (e.g., Kass et al., 2022). Our alphahull and clustered convex hull estimators for boundaries differ because they focus on exterior points in spatial sets. Even so, accurate representation requires a critical minimum number of sightings in areas not advantaged by transport access, high population density, or sufficient disposable income. As the occurrence of sightings in non-advantaged areas increases, so does the accuracy of boundary estimation (Feeley and Silman, 2011). Given that the GBIF occurrence inventory is growing by about 1.3 million new reports per day, one can expect that, over time, increased sightings will improve the boundary estimates for sparsely reported species.
3.1 Pilot example
To illustrate the results of our mapping algorithm exercise, we take the example of the species Lagidium viscacia (common name: Mountain Viscacha), whose range extends from areas in Argentina and Chile to Bolivia and Peru. In Fig. 1, a comparison of panels a and b shows that the reported sightings of this species include a few points beyond the northern boundary of the expert range map (Burgin et al., 2020a), along with many points beyond the southern boundary. Panel c, which displays the output of our mapping algorithm, shows that the alphahull boundary follows the curvilinear north-south orientation of the point set, widening and narrowing as the point set expands and contracts. As shown, it overlaps heavily with the map of Burgin et al. (2020a), but extends its northern and southern boundary areas to incorporate these sightings.
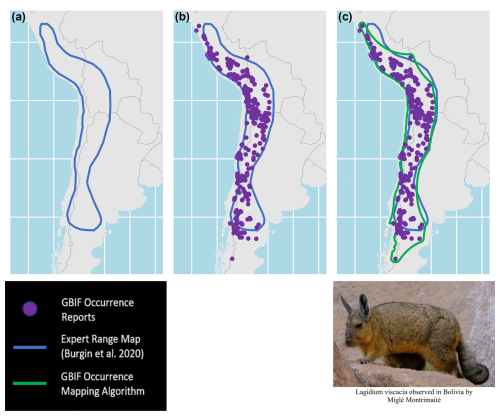
Figure 1Mapping exercise results for Lagidium viscacia (Mountain Viscacha), showing overlapping boundaries: (a) Expert range mapping boundary in blue from Burgin et al. (2020a), (b) Overlay of GBIF occurrence report locations in purple with the expert range mapping boundary in blue from Burgin et al. (2020a), and (c) Overlay of the GBIF occurrence mapping algorithm in green with GBIF occurrence report locations in purple with the expert range mapping boundary in blue from Burgin et al. (2020a).
3.2 Overall mapping results
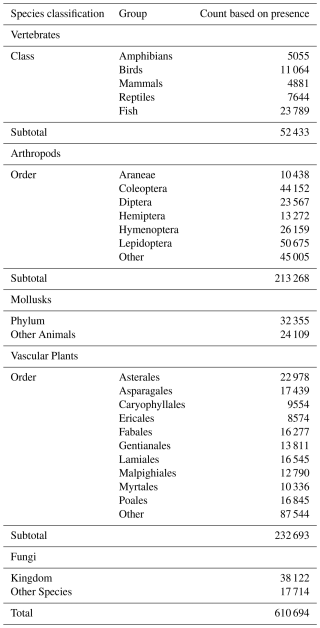
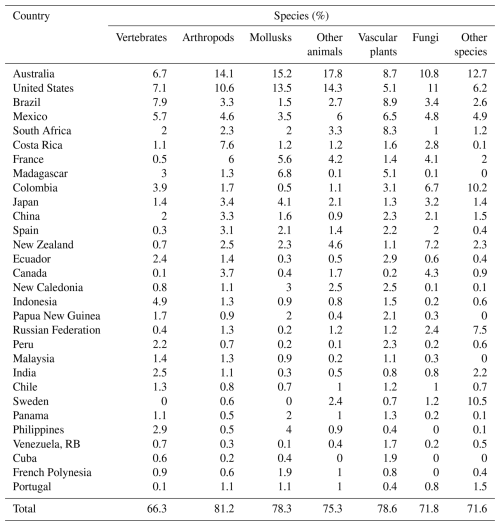
Occurrence record maps were computed for 610 694 GBIF species, comprising 52 433 vertebrates (amphibians, birds, fish, mammals, and reptiles), 213 268 arthropods, 32 355 mollusks, 24 109 other animals, 232 693 plants, 38 122 fungi, and 17 714 other species (the kingdoms Archaea, Bacteria, Chromista, Protozoa, and Viruses) (Table 1).
The exercise was limited to GBIF species in three kingdoms (Animalia, Plantae, and Fungi) with at least three unique geolocated occurrences since 1970. To remove spurious observations from locales (e.g., zoos and botanical gardens), we relied on the following two methods: (1) exclusion of isolated outlier occurrences before map estimation, which happens automatically in our mapping algorithms and (2) exclusion of bounded point sets with fewer than three observations after map estimation. For the many species maps that have multiple bounded areas, we imposed a conservative interpretation of the evidence, dropping species maps with single-bounded areas when they contain fewer than three observations. Although this final condition may seem redundant; it is important to include as a species can pass the initial condition and fail the final one since our estimation algorithms may exclude an outlier point or two from their computations of the bounded areas. Examples of the convex and alpha hull comparisons are included in Appendix A (Sect. 2).
3.3 Case comparisons
To test our species boundary mapping from the current inventory, we ask whether the view of global biodiversity distribution it provides is consistent with that of existing expert maps. On comparing our estimated GBIF occurrence maps with publicly available expert maps from recently published research, we find that thousands of species with GBIF maps have been mapped by the research teams. Using these matched species, each comparison assesses the similarity in global biodiversity patterns produced by our GBIF maps and the expert research products by rank group based on the pixel level distribution of the species maps from 1 (lowest count) to 10 (highest count). Where the patterns diverge, we explore the technical factors that can explain the differences. The first case comparison retains the traditional focus on vertebrates, comparing mammal range maps estimated by Marsh et al. (2022). The second one focuses on a comparison with maps for ants developed by Kass et al. (2022), while the third centers on a limited set of vascular plants mapped by Borgelt et al. (2022). Invertebrates are significantly underrepresented in existing expert maps (Kass et al., 2022). At the outset, it should be noted that this study's major contribution is the expanded coverage of invertebrates. As shown in Table 1, our work offers more comprehensive representation by estimating maps for 213 268 arthropods.
3.3.1 Mammals
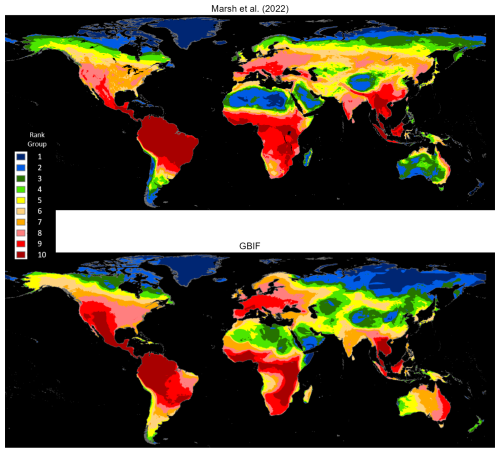
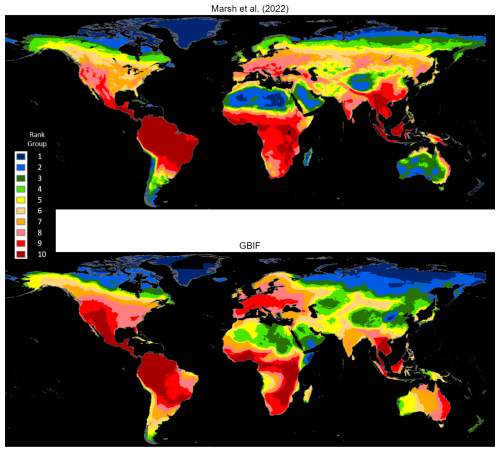
Marsh et al. (2022) map the native ranges of mammals globally using the authoritative taxonomy provided by the Mammal Diversity Database (Burgin et al., 2018). Their exercise harmonizes species maps from the Checklist of the Mammals of the World (Burgin et al., 2020b, c; MDD, 2020), and the Handbook of the Mammals of the World published in nine volumes by Mittermeier et al. (2013), Wilson et al. (2016, 2017), and Wilson and Mittermeier (2009, 2011, 2014, 2015, 2018, 2019). In our GBIF occurrence maps database, we identify 3530 mammals that are also mapped by Marsh et al. (2022). We rasterize both sets of maps using a global grid with 0.05° (5 km) resolution. For each species map, rasterization assigns a value of 1 to grid cells that overlap with the map and 0 to other cells. Next, we compute species densities by cellwise addition across 3530 rasters for each set. Figure 2 compares cell counts, which are ranked in 10 groups. The maps' broad patterns are visibly similar. Both assign ranks in the highest two groups to Central America, northwest South America, West Africa, East Africa, the northern region of Central Africa, the eastern region of southern Africa, Western Europe and Southeast Asia. Notably, they also differ in other regions. The GBIF map assigns higher ranks to large areas of Mexico, western United States, and eastern Australia and lower ranks to the southeastern Amazon region and South Asia (also see Fig. A1).
Technical differences can explain divergences in the two patterns. For example, the Marsh et al. (2022) maps estimate native ranges of global mammals, taking into account recorded historical occurrences and biogeographical factors that correlate with the range of each mammal species. By contrast, the GBIF mammal maps bound the areas where species occurrences have been reported since 1970. Regions where GBIF ranks are higher than Marsh et al. (2022) ranks have many species with reported occurrences beyond their estimated native ranges; regions with lower GBIF ranks have occurrence reports clustered in subareas within native ranges. This difference could reflect underreporting for GBIF species in lower-ranked areas, although many higher-ranked areas appear similarly disadvantaged for species observation. In our view, the more plausible explanation is that lower-ranked regions are populated by many species whose ranges have contracted over time. The ongoing accumulation of GBIF species occurrence reports should help to resolve this issue.
Figure 2 compares 3530 mammals with maps in both databases, while Fig. 3 does the same for all mapped terrestrial mammals (4138 for GBIF and 6360 for Marsh et al., 2022). Comparison with Fig. 2 reveals almost no difference for GBIF; however, Marsh et al. (2022) have generally higher rankings for Indonesia and Papua New Guinea. Mammal species may be underrepresented in GBIF occurrence reports from the two countries, although this seems more likely for sparsely populated Papua New Guinea than densely populated Indonesia (also see Fig. A2). The more likely explanation, in our view, is that the areas populated by many mammals have contracted.
3.3.2 Ants
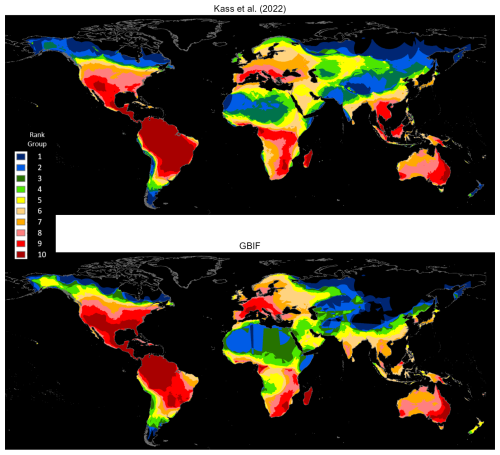
Clark and May (2002) identified a severe taxonomic bias in conservation research, finding that vertebrates accounted for only 3 % of described species but 69 % of published papers. Conversely, invertebrates accounted for 79 % of described species and just 11 % of published papers (Leather et al., 2008). Kass et al. (2022) address this problem for ants using a variety of datasets and techniques, including the alphahull algorithm, to estimate the range maps. In our GBIF occurrence maps database, we identify 5445 ant species also mapped by Kass et al. (2022). We rasterize both sets of maps using a global grid with 0.05° (5 km) resolution and compute species densities by cellwise addition across 5445 rasters for each set.
Figure 4 compares the cell counts, which are ranked in 10 groups. Many areas exhibit similar patterns, including northern North America, Mexico, Central America, northwest South America, Eastern and Western Europe, West Africa, southern Africa, Madagascar, and eastern Australia. However, there are three notable differences. First, both maps identify a large high-ranking region in the Western Hemisphere, which is further north for GBIF than for Kass et al. (2022). Second, both maps identify a band of relatively high ranks across West and northern Central Africa, linking to a north-south band in East and southern Africa; however, the rankings for GBIF are generally lower than those for Kass et al. (2022). Third, Southeast Asia ranks uniformly higher for Kass et al. (2022) than for GBIF (also see Fig. A3).
Since Kass et al. (2022) also rely heavily on the alphahull methodology, we attribute these differences to two technical factors. First, their database comes from intensive processing and error checking of records drawn from the Global Ant Biodiversity Informatics (GABI) database in July 2020. In our study, by contrast, the records are drawn from GBIF occurrence data, as of July 2023. Second, our approach is significantly more conservative. For example, we exclude unique species occurrences that number fewer than three, while Kass et al. (2022) include them; since alphahulls cannot be estimated for these 5168 ant species, Kass et al. (2022) estimate their ranges by drawing 30 km buffer zones around the occurrence locations. Given this difference, comparing full database results for GBIF and Kass et al. (2022) would be, in effect, comparing apples and oranges.
3.3.3 Vascular plants
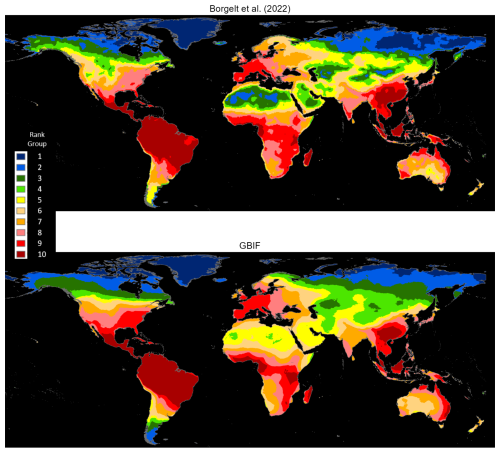
Borgelt et al. (2022) have recently developed spatial density maps for vascular plants in the International Union for Conservation of Nature (IUCN) Red List (IUCN, 2022). They utilize maximum entropy (Maxent) models that predict the likelihood of species occurrences from the values of several environmental variables. For each species, they identify native regions from a web-scraping exercise using the Plants of the World Online (POWO) database, with regional identification standardized from the World Geographical Scheme for Recording Plant Distributions (WGSRPD). Typically, the resulting native regions are the boundaries of small countries or provinces (Database of Global Administrative Areas, GADM, level-1 administrative units) in large countries. Borgelt et al. (2022) estimate the models using GBIF occurrence data with restrictive prior conditions. To preserve compatibility with the environmental variables used for Maxent estimation, the data are confined to the 2000–2020 period. For each species, georeferenced occurrence reports exclude all observations outside pre-identified native regions, and Maxent-estimated species distributions are also confined to native regions. The advantage of this approach is that it guarantees the exclusion of spurious observations from such entities as botanical gardens and private collections in other regions. One drawback, however, is that it incurs the cost of excluding potentially large numbers of occurrence observations that lie outside pre-identified native regions that are arbitrarily defined by national or provincial boundaries.
Unlike Borgelt et al. (2022), our exercise is not constrained by the need for compatibility with environmental modeling variables; therefore, we draw on a longer time period (1970–2023). Also, we impose no prior geographic restrictions on the data. As previously explained, our methodologies estimate occurrence map boundaries after eliminating spurious single outliers and small, isolated occurrence clusters. We identify 32 339 vascular plant species found in both databases, and, as before, rasterize our occurrence maps and compute cell counts at 5 km resolution. As Borgelt et al. (2022) provide species maps in a raster stack with much coarser resolution (50 km), we next extract raster layers for these 32 339 common species and add across layers to obtain relative incidence scores for the 50 km grid cells. Finally, we use mean smoothing to approximate the effect of higher resolution.
Figure 5 displays the comparative results as ranks in 10 groups; the two maps share essentially the same density pattern, except for the somewhat more extensive high-ranking areas in South and Southeast Asia for the Borgelt et al. (2022) maps and the United States for our study's maps (also see Fig. A4).
3.4 Summing up
In all three case comparisons, we find quite similar global patterns of species density with strong agreement between the sources for each case. From a confusion matrix of 5 to 10 classes based on distribution of the species rank group, we find that across these taxa overall accuracy and agreement are significantly better than random (accuracy p-value = 0) with plants consistently outperforming ant and mammals (see discussion in Appendix A1, and Table A1). Additionally, we calculated the local Spearman correlation coefficient for each case and found high agreement in areas with low population and levels of lights at night. Where the patterns diverge, the discrepancies can be traced to technical differences. In the case of mammals, differences between the GBIF and expert native-range maps can be attributed to either undershooting, where expert map boundaries exclude many GBIF occurrences, or overshooting, where GBIF occurrences are persistently absent in parts of the native-range maps. In the case of ants, where the research also utilizes alphahull estimation, differences are attributable to differences in source databases and our relatively conservative approach to map estimation. Finally, in the case of vascular plants, where the research employs GBIF occurrences and the global pattern similarity is most striking, the few discrepancies are attributable to temporal and spatial restrictions imposed by the expert research team.
The effectiveness of biodiversity conservation plans will require identification of occurrence regions for species with elevated extinction risks. While multiple approaches exist for setting conservation priorities, decisions on resource allocation are typically made at national and subnational levels, particularly in the Low- and Middle-Income Countries. Endemic species and those with very restricted ranges provide a practical entry point, as they are highly vulnerable, confined to smaller geographic areas, and relatively straightforward to target with conservation measures. Protecting these species not only contributes to global biodiversity goals but also safeguards the unique natural heritage of individual nations, creating an added incentive for action. Therefore, using the maps developed with GBIF data, we explored (1) species endemic to a single country and (2) species under continuous threat owing to their small occurrence regions. Our database reflects GBIF-sourced occurrence maps for previously unmapped species, as well as revised estimates for those with existing maps.
4.1 Endemic species distribution by group and country
With the new dataset, we explored the endemic status assigned to species that are 100 % resident in a single country. By this criterion, 44.6 % (272 189) of the 610 694 species maps tabulated by country are classified as endemic. The incidence of endemism differs widely by species group (e.g., 54.5 % for mollusks, 48.7 % for vascular plants, 47 % for other animals, 44 % for arthropods, 37.1 % for vertebrates, and 29.5 % for fungi).
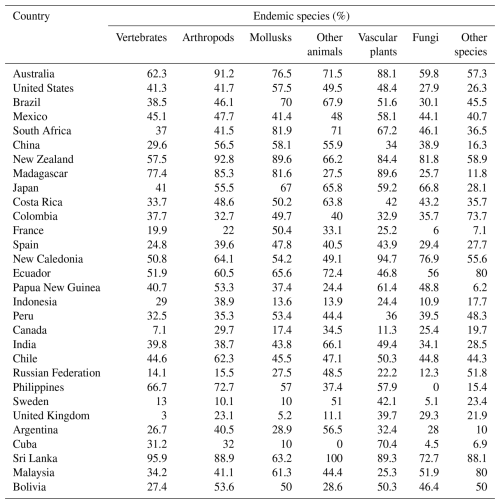
We also count endemic species by country and species group. Country scale plays a major role in raw counts, so we standardize by total country species to highlight the relative importance of endemicity in each country and species group. Table 2 provides a summary for the top 30 countries in each species group, sorted in descending order by average ranking for the seven groups. Overall, the top 30 have 86.6 % (235 706 out of 272 189 species); and our results assign overall top 10 status to Australia, United States, Brazil, Mexico, South Africa, China, New Zealand, Madagascar, Japan, and Costa Rica. Even for the top 30 countries, endemicity varies enormously by species group. In terms of vertebrates, for example, 62 % are endemic in Australia versus only 3 % in the United Kingdom. For plants, the endemicity in Madagascar, Australia, and New Zealand is extremely high, at 89.6 %, 88.1 %, and 84.4 %, respectively. For arthropods, the maximum endemicity is even higher in New Zealand and Australia, at 92.8 % and 91.2 %, respectively. Mollusks, fungi, and other species more closely resemble arthropods and vascular plants in the relative compactness of their ranges.
It should be noted that Table 2 excludes small island territories that rank high in at least one group, including South Georgia, French Polynesia, Heard and McDonald Islands, Norfolk Island, and the Malvinas/Falklands non-determined legal status territory.
4.2 Distribution of small-region species
Small range size has been studied extensively in the empirical literature (Jenkins et al., 2015; Kraus et al., 2023; Manne et al., 1999; Manne and Pimm, 2001; Purvis et al., 2000; Veach et al., 2017). Jenkins et al. (2015), for example, note that “small range size is the best predictor of extinction risk and, thus, the first metric for conservation priority.” It has particular significance since it is a widely recognized indicator of extinction risk that is computable for any species that can be mapped.
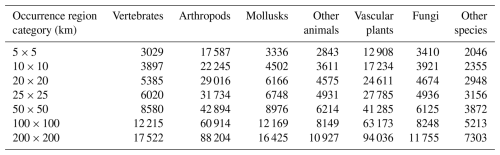
However, it should be noted that small-range status is not determinate; there is no single, critical minimum habitat size, given the myriad interactions between species and habitat characteristics that affect extinction risks. Therefore, we examined the size and global distribution of species with small occurrence regions in our GBIF maps database, considering the effects of changing the criteria for small-occurrence-region status. Table 3 displays the cumulative global count for species groups as the occurrence region increases from 5 km × 5 km to 200 km × 200 km. Even for occurrence regions of 10 km × 10 km or less, 57 765 species are identified; this number increases to 85 310 at 25 km × 25 km or less. Differences across species groups reflect their varying representation in the database and group-specific factors.
We believe that an upper bound of 25 km × 25 km on a critical scale for small-range species is appropriately conservative. The small-range species count increases to 117 946 at 50 km × 50 km or less, 170 081 at 100 km × 100 km or less, and 246 172 at 200 km × 200 km. From a policy perspective, the feasibility and sustainability of species protection tend to decline as the number of species protected increases. Since even the 25 km × 25 km limit qualifies nearly 85 310 species as having a small occurrence region, we retain it here, recognizing that other analyses may well opt for higher limits.
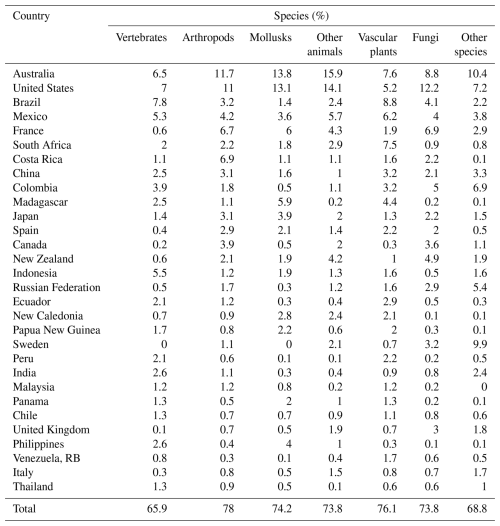
Using GIS overlays of GBIF maps and country boundaries, we count species with small occurrence regions by country, finding that their international distribution is skewed. The top 30 countries account for 75.5 % of them (64 443 out of 85 310 species). Our overall results assign top 10 status to Australia, United States, Brazil, Mexico, France, South Africa, Costa Rica, China, Colombia, and Japan. Australia leads with 8673 species, followed by the United States (7791), Brazil (4434), Mexico (4217), and France (3732). Comparing Table 3 with Table 4 suggests that small-occurrence-region species are endemic in most cases, so the dominant country is chosen by default. In other cases (e.g., Panama, Venezuela, RB, Thailand, and Italy), it is the country with greatest area share in the species' GBIF occurrence map. Among species groups, the top 30 countries' global share varies from 66 % (vertebrates) to 78 % (arthropods) (Table 4).
4.3 Distribution of endemic species with small occurrence regions
We also explored the geographical distribution of endemic species with small occurrence regions (25 km × 25 km size limit). Our results identified 67 941 species in a single country (Table 5).
As before, we find that the international distribution is skewed, with 78.2 % (53 114) of the 67 941 species found in 30 countries. The overall results assign top 10 status to Australia, United States, Brazil, Mexico, South Africa, Costa Rica, France, Madagascar, Colombia, and Japan. Australia leads with by 8072 species, followed by the United States (6003), Brazil (3629), Mexico (3621) and South Africa (2911). Among species groups, the top 30 countries have the following global shares: arthropods (81.2 %), vascular plants (78.6 %), mollusks (78.3 %), other animals (75.3 %), fungi (71.8 %), other non-animal and non-plant species (71.6 %), and vertebrates (66.3 %).
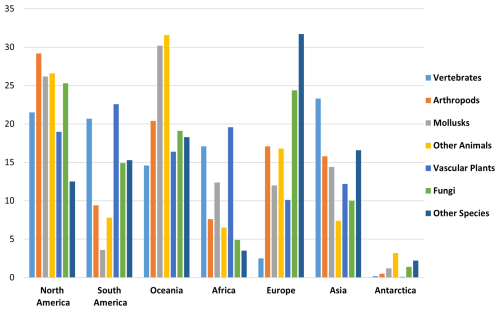
Among endemic species with small occurrence regions, the largest share is found in Oceania (28 %), followed by North America (23 %). Four regions are in the mid-range – South America (15 %), Asia (13 %), Africa (11 %), and Europe (9 %) – and Antarctica has small representation, at 1 % (Fig. 6).
4.4 Candidate hotspot areas for protection
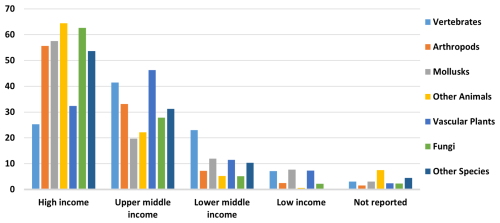
Limited resources for biodiversity conservation make it critical to prioritize protection efforts in regions inhabited by many unique, at-risk species. Endemism and small occurrence regions, as identified by our maps, can inform conservation policy priority-setting. This study's findings indicate that 40 countries have significant opportunities for protecting areas with concentrations of endemic species, species with small occurrence regions, and species with both features. Aligning countries with World Bank income groups reveals an encouraging trend for conservation. While over 4000 endemic species with small ranges are in Low- and Lower-Middle-Income countries, the majority, 82.7 %, are in high and upper-middle-income countries (Fig. 7), which generally have substantial conservation resources. Many such areas may already be protected. Although a global assessment was beyond this study's scope, it would be a valuable future application of our GBIF species maps database.
It should be noted that the maps constructed with processed data also provide opportunities for understanding the geographic distribution of the species within countries.
Transparent and accessible biodiversity data remain central to informed decision-making for conservation and to safeguarding a livable planet for current and future generations. However, comprehensive and all-taxa-inclusive biodiversity data have long been scarce, limiting the ability to assess species distribution and conservation priorities at a global scale. This study presents the first near-comprehensive global mapping of terrestrial, freshwater, and marine biodiversity derived from GBIF occurrence data. By applying the alphahull algorithm and complementary clustering techniques to over 600 000 species, the analysis demonstrates that high-resolution, occurrence-based approaches provide a robust complement to expert-drawn range maps. Building on these results and given the numerous competing demands on land and water, the spatial precision of conservation activities and policies can be enhanced through alphahull techniques, thereby increasing their potential for success by effectively focusing the area of consideration. Also, our effort aligns with that of Janicki et al. (2016), and a salient contribution of this work is the inclusion of previously unmapped taxa – particularly invertebrates, fungi, and plants – which substantially broadens the taxonomic and spatial scope of global biodiversity representation.
5.1 Expanding the biodiversity evidence base
The observed alignment between occurrence-based and expert-derived maps underscores the potential of crowdsourced georeferenced data for large-scale biodiversity mapping. Discrepancies between the two primarily reflect temporal and ecological scope. Expert maps typically represent potential distributions based on habitat suitability and species ecology, while occurrence-based maps capture realized distributions shaped by recent environmental and anthropogenic dynamics (Meyer et al., 2016). Consequently, GBIF-derived boundaries can more effectively detect recent range contractions or expansions associated with deforestation, climate change, or land-use conversion. The rapid updating of GBIF records – currently exceeding one million new georeferenced occurrences daily – creates opportunities for near-real-time monitoring of biodiversity change, a capability unavailable in static expert compilations.
5.2 Addressing taxonomic and geographic bias
A major contribution of this work lies in filling taxonomic gaps that have historically constrained global biodiversity assessments. Existing global datasets emphasize vertebrates, overlooking invertebrates and fungi that together comprise the majority of known species (Cardoso et al., 2011). The integration of over 213 000 arthropods, 38 000 fungi, and 24 000 other animal taxa demonstrates the feasibility of incorporating these groups systematically into global-scale analyses. While GBIF data remain spatially biased toward accessible and high-income regions (Yesson et al., 2007), ongoing growth in citizen science platforms and institutional data sharing is steadily improving geographic representativeness. As observation density increases, particularly across tropical and remote regions, boundary estimation will gain precision, enabling more accurate identification of small-range and data-poor species.
5.3 Implications for conservation priority setting
The identification of endemic and small-range species from GBIF-derived maps provides new insights for conservation targeting. A broader view of biodiversity provides an opportunity for more countries to contribute to a shared vision of conservation stewardship, as most nations host significant distributions of species within at least one major taxonomic group. Consistent with prior studies (Kier et al., 2009), this analysis confirms that small-range species are disproportionately at risk of extinction due to limited dispersal capacity and habitat specialization. The correlation between endemism and small-range occurrence suggests that national-level conservation actions will be critical for achieving global biodiversity outcomes. Australia, the United States, Brazil, Mexico, and South Africa emerge as global centers of endemism and small-occurrence-region species, underscoring their unique biodiversity assets. The identification of numerous localized hotspots – such as Madagascar, Costa Rica, and Southeast Asia – further highlights regions where protection gap analyses are most urgent.
5.4 Policy relevance and the Global Biodiversity Framework
Implementing the goal of protecting 30 % of the planet's land and sea areas by 2030, as set out in Target 3 of the 2022 Kunming–Montreal Global Biodiversity Framework (GBF), requires precise identification of areas critical for conservation. Using occurrence data for over 600 000 species from GBIF, this study provides the largest set of species distribution maps derived from open-access data and highlights 30 countries that could play central roles in area-based conservation, capturing 86.6 % of endemic species, 75.5 % of small-occurrence-region species, and 78.2 % of species that are both endemic and range-limited. These data can also be used to examine the overlap of existing protected areas with endemic-species distributions, highlighting significant variation in initial conservation conditions, including current protection levels and the spatial distribution of unprotected species (Dasgupta et al., 2025a, b).
The GBIF database, expanding by approximately 1.3 million new records daily, allows continuous updates for previously unmapped species and refinement of existing maps. The estimation algorithm supports this growth, providing area estimates for all species and complementing traditional risk indicators. In addition, the alpha hull approach offers a parsimonious, occurrence-driven estimate of species range, requiring minimal assumptions and no environmental predictors. Our practical experience in client countries further indicates that communicating with policymakers is often more straightforward when using species occurrence regions derived from reported occurrences, compared to statistical modeling methods that infer occurrence from environmental predictors, which can be challenging to explain. Given the rate of biodiversity loss, it is urgent to initiate the biodiversity dialogue immediately, beginning with approaches to mapping that are readily understandable. The open-access nature of the data, algorithms, and maps aligns with the GBF's emphasis on equitable sharing, transparency, and evidence-based policy design. High-resolution, taxonomically inclusive maps can guide national and transboundary prioritization, while continuous updating allows adaptive policy responses – an essential feature for tracking progress toward GBF targets and ensuring accountability in biodiversity monitoring frameworks. This need for spatially explicit and scalable information is particularly relevant for international institutions and development organizations. An information paradox often confronts decision-makers in transnational institutions: while the scope of their operations is global, the scale of specific interventions is inevitably local or regional (Blankespoor et al., 2023).
Looking ahead, open and equitable sharing of biodiversity, threat, and protection data across administrative boundaries remains essential, as species and ecosystems frequently span political jurisdictions (Blankespoor et al., 2025; Dasgupta et al., 2025c; Tanalgo, 2025). Reliable, high-resolution, and up-to-date data facilitate coordinated conservation strategies across terrestrial, freshwater, and marine systems. Healthy biodiversity underpins ecosystem stability and sustains essential services such as agriculture, fisheries, water security, and climate resilience. It is therefore fundamental to sustainable development, poverty alleviation, equitable prosperity, and safeguarding a livable planet for current and future generations.
5.5 Limitations and future directions
Despite these advances, several limitations warrant acknowledgment. First, GBIF occurrence data remain uneven in spatial and temporal coverage, with collection bias favoring regions with established research infrastructure, accessibility and funding (Beck et al., 2014; Boakes et al., 2010; Hickisch et al., 2019; Faxon and Chapman, 2025). Hortal et al. (2015) discuss shortfalls of biodiversity data including taxonomic (Linnean), distribution (Wallacean), and abundance (Prestonian). First, taxonomic challenges can persist because research effort is often linked to a disproportionate concentration on early-described taxa, as in the case of 10 000 reptile species examined by Guedes et al. (2023). In a study of the Eastern Arc Mountains in Tanzania, Ahrends et al. (2011) find suggestive evidence of biodiversity patterns linked with investment in inventories. Second, accurate species occurrence records are needed to improve the understanding of species distribution mitigating the “Wallacean Shortfall.” Third, presence-only data do not capture species abundance, which limits inference about population viability within mapped boundaries. Fourth, while the alphahull method provides an efficient means of delineating occurrence extents, it does not explicitly account for environmental suitability or ecological connectivity. Future research could enhance these methods by integrating species distribution modeling, remote-sensing covariates, sampling, latitudinal taxonomic gradient and temporal updating to capture dynamic species–environment interactions (Elith and Leathwick, 2009; Diniz-Filho et al., 2023).
Although GBIF strengthens the data ecosystem providing a common repository for data collections from scientists, agencies, community science and museums (Heberling et al., 2021), further investment would help improve GBIF completeness by building local digitization and curation capacity and by integrating diverse data streams. Empirical work in the Amazon illustrates both the promise and challenges of current approaches: de Araujo et al. (2022) find that combining global and national databases can increase biodiversity knowledge and reduce inventory gaps. Sustained efforts to digitize, publish, and improve data quality are essential for keeping biodiversity data efficient, effective, and relevant (Ball-Damerow et al., 2019). Targeted capacity-building programs, such as GBIF's Biodiversity Information for Development (BID), support data mobilization, training, and publisher engagement to improve the coverage and usability of biodiversity data across the world (GBIF, 2025).
These data are available at the World Bank's Development Data Hub under Global Biodiversity Species Occurrence Gridded Data and Global Biodiversity Species Occurrence Endemism and Small Range Data. The datasets can be accessed at https://doi.org/10.57966/h21e-vq42 (Dasgupta et al., 2024). The authors utilized Google BigQuery to retrieve species occurrence records and R software for data processing. The associated scripts are available from the authors upon request. Appendix B includes a flowchart of the main processing algorithm.
Overall, this study demonstrates the feasibility and scientific value of automated, open-access biodiversity mapping at global scale. By complementing expert-based datasets with data-driven, continuously updatable occurrence maps, it bridges long-standing taxonomic and geographic data gaps, improves spatial precision for conservation planning, and enhances the policy relevance of biodiversity information systems. As global data sharing and digital observation platforms continue to expand, the approach outlined here can provide a scalable and transparent foundation for evidence-based implementation of the Global Biodiversity Framework and related multilateral environmental agreements.
A1 Case Comparisons of Species Occurrence Regions and expert maps
To test our species boundary mapping from the current inventory, we compare the global biodiversity patterns derived from our current inventory with those depicted in expert-generated maps. By aligning our GBIF-based occurrence maps with expert maps from recent studies, we identify thousands of overlapping species. These shared species serve as the basis for assessing how closely our GBIF-derived biodiversity patterns match those reported in the expert research.
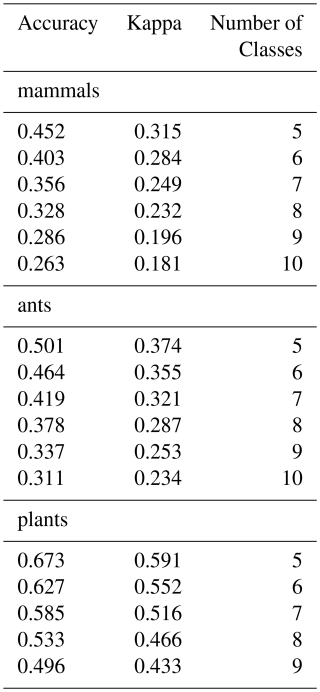
We compare our Species Occurrence Regions derived from GBIF with the sets of matched species for mammals (Marsh et al., 2022), ants (Kass et al., 2022) and plants (Borgelt et al., 2022) with a confusion matrix with 5 to 10 classes by distribution (see Table A1). Model performance varied across taxa and the number of classes. For mammals, overall accuracy was highest at 0.452 (κ = 0.315) with 5 classes to the lowest at 0.263 (κ = 0.181) with 10 classes, indicating that model agreement beyond chance declined as class complexity increased. Ants showed slightly higher performance at low class counts, with accuracy of 0.501 (κ = 0.374) for 5 classes, dropping to 0.311 (κ = 0.234) for 10 classes. Plants were the best-predicted group, achieving 0.673 accuracy (κ = 0.591) for 5 classes and 0.450 accuracy (κ = 0.389) for 10 classes. Across all taxa, accuracy confidence intervals were narrow, reflecting precise estimates, and all comparisons were significantly better than random (Accuracy p-value = 0). These results suggest that predictive performance declines with increasing class granularity, with plants consistently outperforming ants and mammals in overall accuracy and agreement.
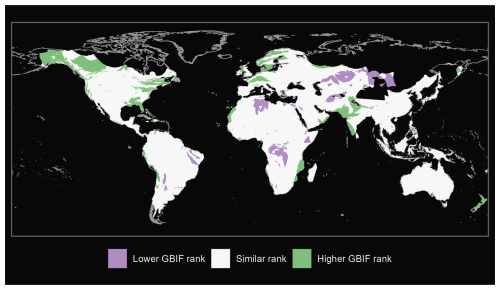
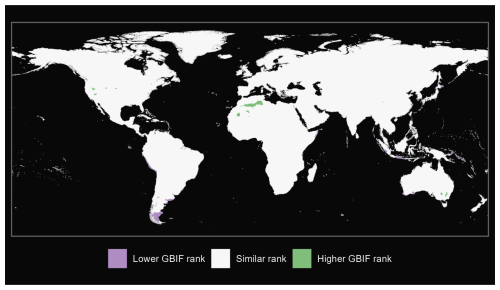
We also produce maps of the differences in ranks between our GBIF-based occurrence maps with the expert maps. We illustrate three categories: (i) GBIF has a lower rank than expert map by more than 2 ranks, (ii) similar rank that includes an absolute difference of 2 ranks or fewer and (iii) GBIF has a higher rank than the expert map by more than 2 ranks (see Fig. A1 for matched mammals, Fig. A2 for full mammals, Fig. A3 for plants and Fig. A4 for ants).
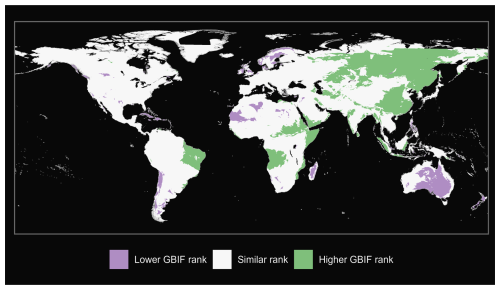
Figure A1Comparison of matched mammals from our GBIF species occurrence regions and expert-based map (Marsh et al., 2022).
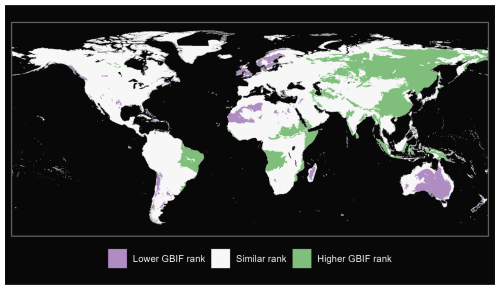
Figure A2Comparison of full mammals from our GBIF species occurrence regions and expert-based map (Marsh et al., 2022).
A2 Case Comparisons of Species Occurrence Regions and expert maps
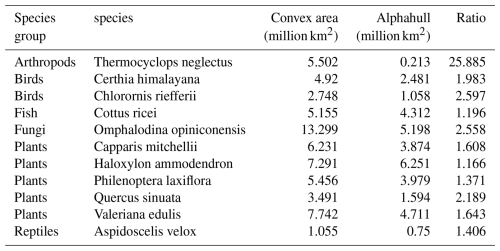
To evaluate differences between spatial modeling approaches, we compared species occurrence regions generated using the convex hull and alpha hull methods across representative taxa, including arthropods, birds, fungi, fish, plants, and reptiles. The convex hull method defines a minimal polygon encompassing all occurrence points, while the alpha hull identifies finer boundaries that can separate spatially distinct clusters (see Sect. 2.2). Table A2 provides the area for each method and the ratio of convex hull area over alpha hull area. Species groups are discussed below with maps of selected species.
A2.1 Arthropods
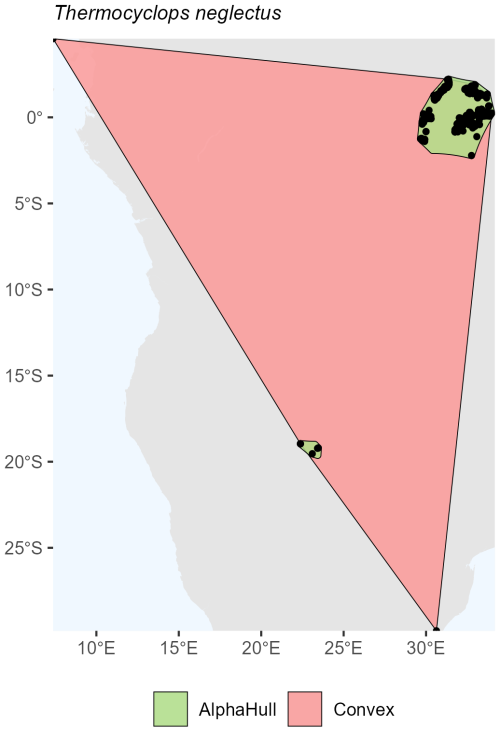
For arthropods, we examined Thermocyclops neglectus, a species broadly distributed across Africa. The convex hull method delineated a large area of approximately 5.50 million km2, encompassing the full extent of known occurrences. In contrast, the alpha hull produced two distinct clusters totaling 0.213 million km2, representing only about 3 % of the convex hull extent (ratio = 25.89). This result illustrates how the alpha hull approach captures core occurrence zones more precisely while excluding large intervening areas of apparent absence (Fig. A5).
A2.2 Birds
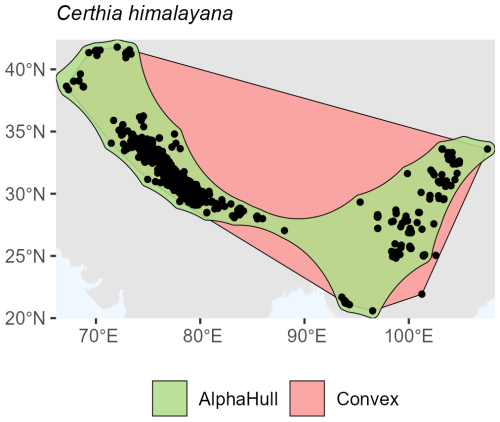
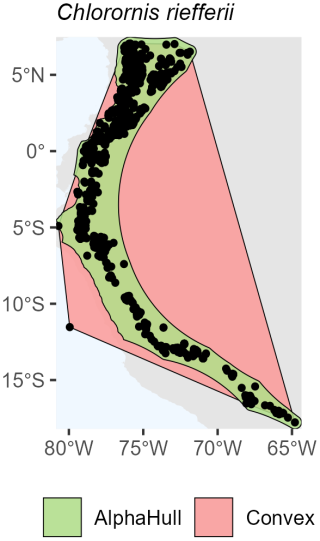
Among birds, we selected Certhia himalayana and Chlorornis riefferii, which represent species with relatively continuous and moderately fragmented distributions, respectively. For Certhia himalayana found in Asia, the convex hull area (4.9 million km2) was about twice as large as the alpha hull area (2.5 million km2), yielding a ratio of 1.98, indicating moderate spatial refinement by the alpha hull (Fig. A6). Chlorornis riefferii found in South America showed a slightly higher ratio (2.60), suggesting more fragmented or discontinuous occurrence points, where the alpha hull excluded peripheral areas of low density (Fig. A7).
A2.3 Fish
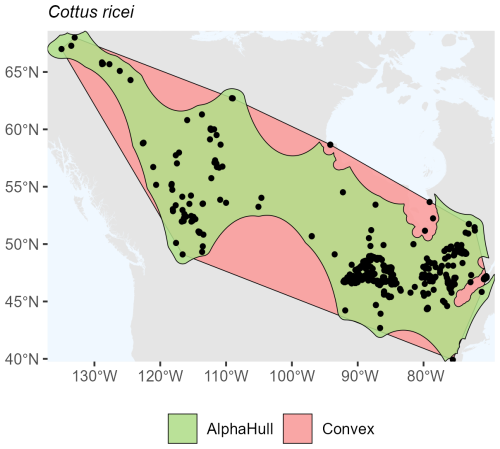
For the freshwater fish Cottus ricei found in North America, the convex hull encompassed an area of 5.16 million km2, only modestly larger than the alpha hull estimate (4.31 million km2, ratio 1.20) (Fig. A8). This relatively low ratio indicates a spatially cohesive set of occurrence records, with both hull methodsapturing a similar extent of distribution.
A2.4 Fungi
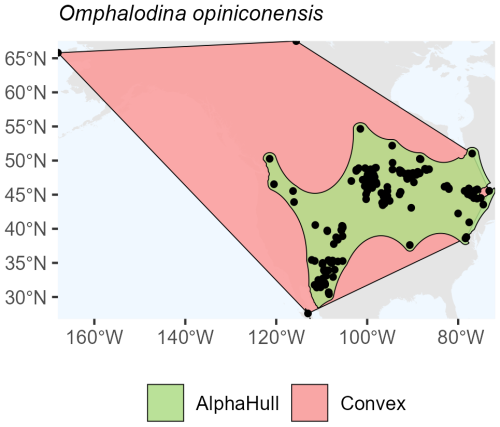
In fungi, the species occurrence regions of Omphalodina opiniconensis show a broad North American distribution, with the convex hull covering 13.3 million km2 and the alpha hull reducing this to 5.2 million km2 (ratio 2.56) (Fig. A9). This pattern suggests that while the species is widespread, its occurrences form distinct spatial clusters that the alpha hull captures more precisely, excluding large unoccupied areas within the convex hull boundary.
A2.5 Plants
Five plant species were analyzed, showing variable differences between the two methods.
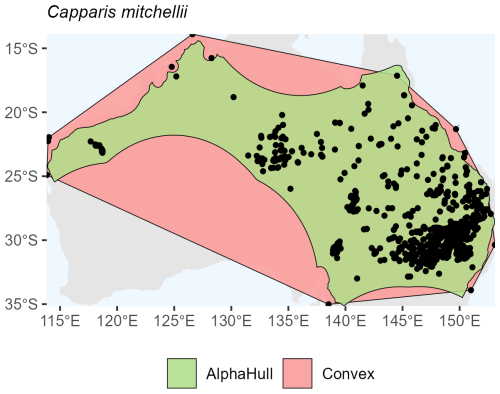
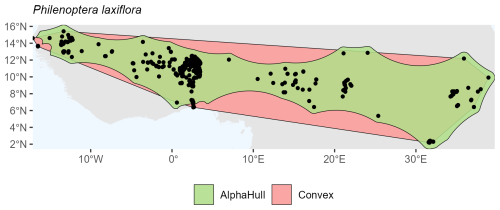
Capparis mitchellii, which is native to Australia, (ratio 1.61) and Philenoptera laxiflora, which is found across Africa, (ratio 1.37) both exhibited moderate spatial contraction under the alpha hull, indicating relatively continuous distributions with some peripheral refinement (Figs. A10 and A11, respectively).
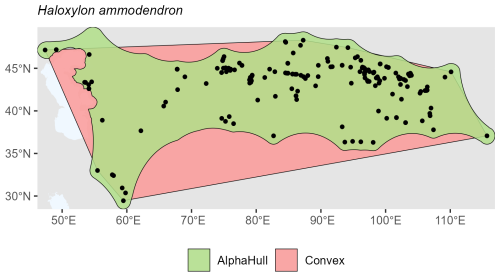
Haloxylon ammodendron, which is found in Asia, showed minimal difference (ratio 1.17), suggesting a dense and cohesive set of occurrences across its range (Fig. A12).
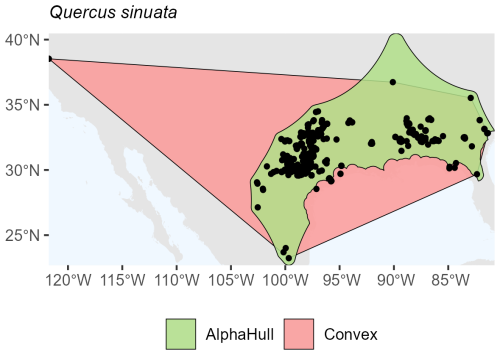
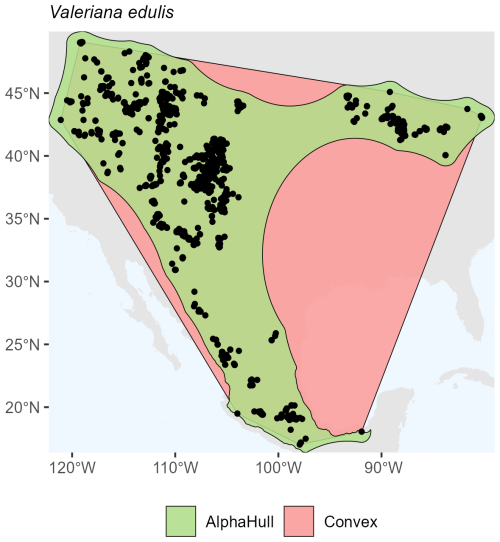
Quercus sinuata, which is found in North America (ratio 2.19) (Fig. A13), and Valeriana edulis, also found in North America (ratio 1.64), had higher ratios, pointing to patchier occurrence patterns or geographic discontinuities (Fig. A14).
Overall, the plant species showed consistent trends where the alpha hull excluded peripheral gaps, providing a more ecologically realistic boundary.
A2.6 Reptiles
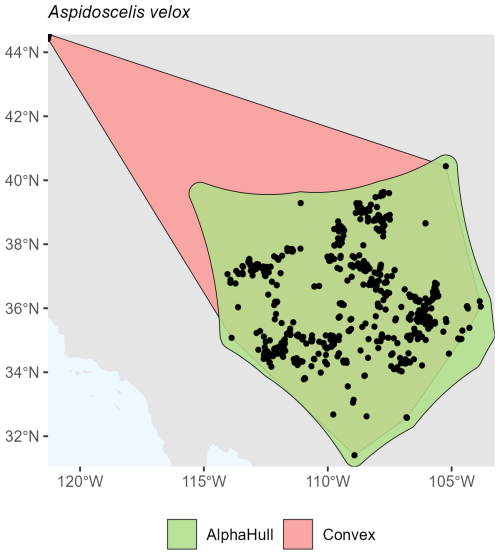
For reptiles, the species Aspidoscelis velox, which is found in North America, showed a convex hull area of 1.06 million km2compared to 0.75 million km2 from the alpha hull (ratio 1.41) (Fig. A15). This moderate difference reflects a relatively compact but slightly fragmented distribution.
Across all taxa, convex hull areas consistently exceeded those derived from alpha hulls, as expected from their respective geometric definitions. The convex hull to alphahull area ratios ranged from 1.17 (H. ammodendron) to 25.89 (T. neglectus), reflecting the degree of spatial clustering among occurrence records. The extreme difference observed for T. neglectus highlights the alpha hull's ability to delineate discrete occurrence clusters and avoid overestimation of occurrence region. Collectively, these findings demonstrate that while convex hulls provide rapid, broad-range approximations, alpha hulls yield finer, more ecologically meaningful representations of species distributions, particularly for taxa with spatially discontinuous occurrences.
From a policy perspective, the results highlight a pattern where convex hull produces areas that are much larger than where the clusters of species occurrences are located. Since there are many competing demands for land and water, the potential success of conservation activities/policy are more precise (spatially) using the alphahull techniques as it focuses the area of consideration.
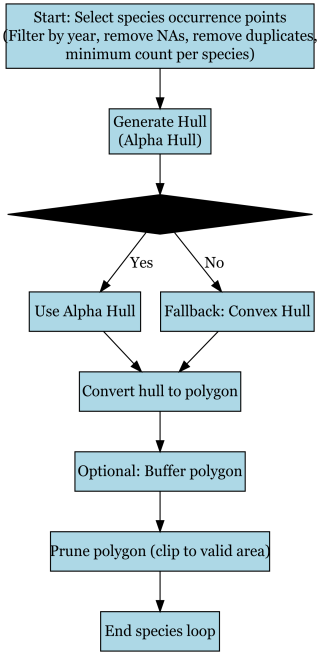
The workflow for species range mapping shown in Fig. B1 begins by selecting species occurrence points, applying a filtering step to remove records with missing coordinates, duplicate entries, or those outside the target time range, while also enforcing a minimum record count per species. After filtering, an alpha hull is generated to approximate each species' distribution. If the alpha hull construction is successful, the result is retained; if not, the procedure falls back to a convex hull. The resulting hull is then converted into a polygon, optionally buffered to smooth edges, and pruned to ensure it aligns with valid geographic boundaries. The process concludes with the finalized range polygon for each species before looping to the next in the dataset.
SD: Conceptualization, Funding Acquisition, Project Supervision, and Writing original draft.
BB: Funding Acquisition; GIS Analysis; Visualization; and Writing, Review, and Editing.
DW: Conceptualization, Data Curation, Methodology, and Formal Analysis.
The contact author has declared that none of the authors has any competing interests.
The findings, interpretations, and conclusions expressed in this article are entirely those of the author(s). They do not necessarily reflect the views of The World Bank, its Board of Executive Directors, or the governments they represent. Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. The authors bear the ultimate responsibility for providing appropriate place names. Views expressed in the text are those of the authors and do not necessarily reflect the views of the publisher.
We are thankful to the World Bank Sustainable Development Global Practice and Environment, Natural Resources and Blue Economy Global Practice for their review and comments. We are also grateful to Polly Means for the graphics and Norma Adams for editorial support.
This research was funded by a grant from the Global Environment Facility (grant no. P179309) to a World Bank program managed by the authors with Dr. Nagaraja Harshadeep Rao. Additional support is from the Space2Stats Program, supported by a grant managed by Brian Blankespoor from the World Bank's Global Data Facility.
This paper was edited by Chaoqun Lu and reviewed by Kenneth Chomitz and two anonymous referees.
Ahrends, A., Burgess, N. D., Gereau, R. E., Marchant, R., Bulling, M. T., Lovett, J. C., Platts, P. J., Wilkins Kindemba, V., Owen, N., Fanning, E., and Rahbek, C.: Funding begets biodiversity, Diversity and Distributions, 17, 191–200, https://doi.org/10.1111/j.1472-4642.2010.00737.x, 2011.
Ball-Damerow, J. E., Brenskelle, L., Barve, N., Soltis, P. S., Sierwald, P., Bieler, R., LaFrance, R., Ariño, A. H., and Guralnick, R. P.: Research applications of primary biodiversity databases in the digital age, PLoS ONE, 14, e0215794, https://doi.org/10.1371/journal.pone.0215794, 2019.
Beck, J., Böller, M., Erhardt, A., and Schwanghart, W.: Spatial bias in the GBIF database and its effect on modeling species' geographic distributions, Ecological Informatics, 19, 10–15, https://doi.org/10.1016/j.ecoinf.2013.11.002, 2014.
Blankespoor, B., Dasgupta, S., Wheeler, D., Jeuken, A., van Ginkel, K., Hill, K., and Hirschfeld, D.: Linking sea-level research with local planning and adaptation needs, Nature Climate Change, 13, 760–763, https://doi.org/10.1038/s41558-023-01749-7, 2023.
Blankespoor, B., Dasgupta, S., and Wheeler, D.: Bridging conflicts and biodiversity protection: the critical role of reliable and comparable data, World Bank Policy Research Working Paper 11076, 32 pp., https://documents.worldbank.org/en/publication/documents-reports/documentdetail/099415202272518792 (last access: 29 October 2025), 2025.
Boakes, E. H., McGowan, P. J. K., Fuller, R. A., Chang-qing, D., Clark, N. E., O'Connor, K., and Mace, G. M.: Distorted views of biodiversity: spatial and temporal bias in species occurrence data, PLoS Biology, 8, e1000385, https://doi.org/10.1371/journal.pbio.1000385, 2010.
Borgelt, J., Sicacha-Parada, J., Skarpaas, O., and Verones, F.: Native range estimates for red-listed vascular plants, Scientific Data, 9, 117, https://doi.org/10.1038/s41597-022-01233-5, 2022.
Burgin, C., Colella, J., Kahn, P., and Upham, N.: How many species of mammals are there?, Journal of Mammalogy, 99, 1–14, https://doi.org/10.1093/jmammal/gyx147, 2018.
Burgin, C., Wilson, D., Mittermeier, R., Rylands, A., Lacher, T., and Sechrest, W.: Illustrated Checklist of the Mammals of the World, Lynx Nature Books, Barcelona, ISBN 978-84-16728-36-7, 2020a.
Burgin, C., Wilson, D., Mittermeier, R., Rylands, A., Lacher, T., and Sechrest, W.: Illustrated Checklist of the Mammals of the World, Vol. 2: Eulipotyphla to Carnivora, Lynx Edicions, Barcelona, ISBN 978-84-16728-36-7, 2020b.
Burgin, C., Wilson, D., Mittermeier, R., Rylands, A., Lacher, T., and Sechrest, W.: Illustrated Checklist of the Mammals of the World, Vol. 1: Monotremata to Rodentia, Lynx Edicions, Barcelona, ISBN 978-84-16728-36-7, 2020c.
Cardoso, P., Erwin, T. L., Borges, P. A. V., and New, T. R.: The seven impediments in invertebrate conservation and how to overcome them, Biological Conservation, 144, 2647–2655, https://doi.org/10.1016/j.biocon.2011.07.024, 2011.
Clark, J. and May, R.: Taxonomic bias in conservation research, Science, 297, 191–192, https://doi.org/10.1126/science.297.5579.191b, 2002.
Dasgupta, S., Blankespoor, B., and Wheeler, D.: Global Biodiversity Data, World Bank Group [data set], https://doi.org/10.57966/h21e-vq42, 2024.
Dasgupta, S., Blankespoor, B., and Wheeler, D.: Country-level pathways to 30 × 30 and their implications for global biodiversity protection, World Bank Policy Research Working Paper 11269, https://documents.worldbank.org/curated/en/099328112032534200 (last access: 25 January 2026), 2025a.
Dasgupta, S., Wheeler, D., and Blankespoor, B.: Pathways to 30 × 30: Evidence-based lessons from global case studies in biodiversity conservation, Diversity, 17, 401, https://doi.org/10.3390/d17060401, 2025b.
Dasgupta, S., Blankespoor, B., and Wheeler, D.: Predicting species extinction risks using occurrence data from the Global Biodiversity Information Facility, Journal of Management and Sustainability, 15, 93–105, https://doi.org/10.5539/jms.v15n2p93, 2025c.
de Araujo, M. L., Quaresma, A. C., and Ramos, F. N.: GBIF information is not enough: national database improves the inventory completeness of Amazonian epiphytes, Biodiversity and Conservation, 31, 2797–2815, https://doi.org/10.1007/s10531-022-02458-x, 2022.
Diniz-Filho, J. A. F., Jardim, L., Guedes, J. J., Meyer, L., Stropp, J., Frateles, L. E. F., and Hortal, J.: Macroecological links between the Linnean, Wallacean, and Darwinian shortfalls, Frontiers of Biogeography, 15, e59566, https://doi.org/10.21425/F5FBG59566, 2023.
Elith, J. and Leathwick, J. R.: Species distribution models: ecological explanation and prediction across space and time, Annual Review of Ecology, Evolution, and Systematics, 40, 677–697, https://doi.org/10.1146/annurev.ecolsys.110308.120159, 2009.
Faxon, H. and Chapman, M.: Beyond spatial bias: understanding the colonial legacies and contemporary social forces shaping biodiversity data, Environmental Research Letters, 20, 064053, https://doi.org/10.1088/1748-9326/add6b6, 2025.
Feeley, K. and Silman, M.: Keep collecting: accurate species distribution modelling requires more collections than previously thought, Diversity and Distributions, 17, 1132–1140, https://doi.org/10.1111/j.1472-4642.2011.00813.x, 2011.
Garcia-Roselló, E., Gonzalez-Dacosta, J., Guisande, C., and Lobo, J.: GBIF falls short of providing a representative picture of the global distribution of insects, Systematic Entomology, 48, 489–497, https://doi.org/10.1111/syen.12589, 2023.
GBIF: GBIF Occurrence, https://console.cloud.google.com/bigquery (last access: 17 February 2024), 2024.
GBIF: BID – Biodiversity Information for Development, https://www.gbif.org/programme/82243/bid-biodiversity-information-for-development (last access: 8 December 2025), 2025.
Guedes, J. J., Moura, M. R., and Diniz-Filho, J. A. F.: Species out of sight: elucidating the determinants of research effort in global reptiles, Ecography, 46, e06491, https://doi.org/10.1111/ecog.06491, 2023.
Guo, W.-Y., Serra-Díaz, J. M., Schrodt, F., Eiserhardt, W. L., Maitner, B. S., Merow, C., Violle, C., Anand, M., Belluau, M., Bruun, H. H., Byun, C., Catford, J. A., Cerabolini, B. E. L., Chacón-Madrigal, E., Ciccarelli, D., Cornelissen, J. H. C., Dang-Le, A. T., de Frutos, A., Dias, A. S., Giroldo, A. B., Guo, K., Gutiérrez, A. G., Hattingh, W., He, T., Hietz, P., Hough-Snee, N., Jansen, S., Kattge, J., Klein, T., Komac, B., Kraft, N. J. B., Kramer, K., Lavorel, S., Lusk, C. H., Martin, A. R., Mencuccini, M., Michaletz, S. T., Minden, V., Mori, A. S., Niinemets, Ü., Onoda, Y., Peñuelas, J., Pillar, V. D., Pisek, J., Robroek, B. J. M., Schamp, B., Slot, M., Sosinski, Ê. E., Soudzilovskaia, N. A., Thiffault, N., van Bodegom, P., van der Plas, F., Wright, I. J., Xu, W.-B., Zheng, J., Enquist, B. J., and Svenning, J.-C: High exposure of global tree diversity to human pressure, Proceedings of the National Academy of Sciences of the United States of America, 119, e202673311, https://doi.org/10.1073/pnas.2026733119, 2022.
Heberling, J. M., Miller, J. T., Noesgaard, D., Weingart, S. B., and Schigel, D.: Data integration enables global biodiversity synthesis, Proceedings of the National Academy of Sciences of the United States of America, 118, e2018093118, https://doi.org/10.1073/pnas.2018093118, 2021.
Hickisch, R., Hodgetts, T., Johnson, P. J., Sillero-Zubiri, C., Tockner, K., and Macdonald, D. W.: Effects of publication bias on conservation planning, Conservation Biology, 33, 1151–1163, https://doi.org/10.1111/cobi.13326, 2019.
Hortal, J., De Bello, F., Diniz-Filho, J. A. F., Lewinsohn, T. M., Lobo, J. M., and Ladle, R. J.: Seven shortfalls that beset large-scale knowledge of biodiversity, Annual Review of Ecology, Evolution, and Systematics, 46, 523–549, https://doi.org/10.1146/annurev-ecolsys-112414-054400, 2015.
Hughes, A. C., Dorey, J. B., Bossert, S., Qiao, H., and Orr, M. C.: Big data, big problems? How to circumvent problems in biodiversity mapping and ensure meaningful results, Ecography, 47, e07115, https://doi.org/10.1111/ecog.07115, 2024.
Isaac, N. and Pocock, M.: Bias and information in biological records, Biological Journal of the Linnean Society, 115, 522–531, https://doi.org/10.1111/bij.12532, 2015.
IUCN (International Union for Conservation of Nature): The IUCN Red List of Threatened Species, https://hosted-datasets.gbif.org/datasets/iucn/iucn-2022-1.zip (last access: 12 March 2024), Version 2022-1, https://doi.org/10.15468/0qnb58, 2022.
Janicki, J., Narula, N., Ziegler, M., Guénard, B., and Economo, E. P.: Visualizing and interacting with large-volume biodiversity data using client–server web-mapping applications: the design and implementation of antmaps.org, Ecological Informatics, 32, 185–193, https://doi.org/10.1016/j.ecoinf.2016.02.006, 2016.
Jenkins, C., Van Houtan, K., Pimm, S., and Sexton, J.: US protected lands mismatch biodiversity priorities, Proceedings of the National Academy of Sciences of the United States of America, 112, 5081–5086, https://doi.org/10.1073/pnas.1418034112, 2015.
Kass, J. M., Guénard, B., Dudley, K. L., Jenkins, C. N., Azuma, F., Fisher, B. L., Parr, C. L., Gibb, H., Longino, J. T., Ward, P. S., Chao, A., Lubertazzi, D., Weiser, M., Jetz, W., Guralnick, R., Blatrix, R., Des Lauriers, J., Donoso, D. A., Georgiadis, C., Gomez, K., Hawkes, P. G., Johnson, R. A., Lattke, J. E., MacGown, J. A., Mackay, W., Robson, S., Sanders, N. J., Dunn, R. R., and Economo, E. P.: The global distribution of known and undiscovered ant biodiversity, Science Advances, 8, eabp9908, https://doi.org/10.1126/sciadv.abp9908, 2022.
Kier, G., Kreft, H., Lee, T. M., Jetz, W., Ibisch, P. L., Nowicki, C., Mutke, J., and Barthlott, W.: A global assessment of endemism and species richness across island and mainland regions, Proceedings of the National Academy of Sciences of the United States of America, 106, 9322–9327, https://doi.org/10.1073/pnas.0810306106, 2009.
Kraus, D., Enns, A., Hebb, A., Murphy, S., Drake, D. A. R., and Bennett, B.: Prioritizing nationally endemic species for conservation, Conservation Science and Practice, 5, e12845, https://doi.org/10.1111/csp2.12845, 2023.
Leather, S., Basset, Y., and Hawkins, B.: Insect conservation: finding the way forward, Insect Conservation and Diversity, 1, 67–69, https://doi.org/10.1111/j.1752-4598.2007.00005.x, 2008.
Mammal Diversity Database (MDD): Mammal Diversity Database (Version 1.2), Zenodo [data set], https://doi.org/10.5281/zenodo.4139818, 2020.
Manne, L. and Pimm, S.: Beyond eight forms of rarity: which species are threatened and which will be next, Animal Conservation, 4, 221–229, https://doi.org/10.1017/S1367943001001263, 2001.
Manne, L., Brooks, T., and Pimm, S.: Relative risk of extinction of passerine birds on continents and islands, Nature, 399, 258–261, https://doi.org/10.1038/20436, 1999.
Marsh, C. J., Sica, Y. V., Burgin, C. J., Dorman, W. A., Anderson, R. C., del Toro Mijares, I., Vigneron, J. G., Barve, V., Dombrowik, V. L., Duong, M., Guralnick, R., Hart, J. A., Maypole, J. K., McCall, K., Ranipeta, A., Schuerkmann, A., Torselli, M. A., Lacher Jr., T., Mittermeier, R. A., Rylands, A. B., Sechrest, W., Wilson, D. E., Abba, A. M., Aguirre, L. F., Arroyo-Cabrales, J., Astúa, D., Baker, A. M., Braulik, G., Braun, J. K., Brito, J., Busher, P. E., Burneo, S. F., Camacho, M. A., Cavallini, P., de Almeida Chiquito, E., Cook, J. A., Cserkész, T., Csorba, G., Cuéllar Soto, E., da Cunha Tavares, V., Davenport, T. R. B., Deméré, T., Denys, C., Dickman, C. R., Eldridge, M. D. B., Fernandez-Duque, E., Francis, C. M., Frankham, G., Franklin, W. L., Freitas, T., Friend, J. A., Gadsby, E. L., Garbino, G. S. T., Gaubert, P., Giannini, N., Giarla, T., Gilchrist, J. S., Gongora, J., Goodman, S. M., Gursky-Doyen, S., Hackländer, K., Hafner, M. S., Hawkins, M., Helgen, K. M., Heritage, S., Hinckley, A., Hintsche, S., Holden, M., Holekamp, K. E., Honeycutt, R. L., Huffman, B. A., Humle, T., Hutterer, R., Ibáñez Ulargui, C., Jackson, S. M., Janecka, J., Janecka, M., Jenkins, P., Juškaitis, R., Juste, J., Kays, R., Kilpatrick, C. W., Kingston, T., Koprowski, J. L., Kryštufek, B., Lavery, T., Lee Jr., T. E., Leite, Y. L. R., Novaes, R. L. M., Lim, B. K., Lissovsky, A., López-Antoñanzas, R., López-Baucells, A., MacLeod, C. D., Maisels, F. G., Mares, M. A., Marsh, H., Mattioli, S., Meijaard, E., Monadjem, A., Morton, F. B., Musser, G., Nadler, T., Norris, R. W., Ojeda, A., Ordóñez-Garza, N., Pardiñas, U. F. J., Patterson, B. D., Pavan, A., Pennay, M., Pereira, C., Prado, J., Queiroz, H. L., Richardson, M., Riley, E. P., Rossiter, S. J., Rubenstein, D. I., Ruelas, D., Salazar-Bravo, J., Schai-Braun, S., Schank, C. J., Schwitzer, C., Sheeran, L. K., Shekelle, M., Shenbrot, G., Soisook, P., Solari, S., Southgate, R., Superina, M., Taber, A. B., Talebi, M., Taylor, P., Vu Dinh, T., Ting, N., Tirira, D. G., Tsang, S., Turvey, S. T., Valdez, R., Van Cakenberghe, V., Veron, G., Wallis, J., Wells, R., Whittaker, D., Williamson, E. A., Wittemyer, G., Woinarski, J., Zinner, D., Upham, N. S., and Jetz, W.: Expert range maps of global mammal distributions harmonized to three taxonomic authorities, Journal of Biogeography, 49, 979–992, https://doi.org/10.1111/jbi.14330, 2022.
Meyer, C., Weigelt, P., and Kreft, H.: Multidimensional biases, gaps and uncertainties in global plant occurrence information, Ecology Letters, 19, 992–1006, https://doi.org/10.1111/ele.12624, 2016.
Mittermeier, R., Rylands, A., and Wilson, D.: Handbook of the Mammals of the World, Vol. 3: Primates, Lynx Edicions, Barcelona, ISBN 978-84-96553-89-7, 2013.
Pateiro-López, B. and Rodríguez-Casal, A.: Generalizing the convex hull of a sample: the R package alphahull, Journal of Statistical Software, 34, https://doi.org/10.18637/jss.v034.i05, 2010.
Pimm, S., Jenkins, C., Abell, R., Brooks, T., Gittleman, J., Joppa, L., Raven, P., Roberts, C., and Sexton, J.: The biodiversity of species and their rates of extinction, distribution, and protection, Science, 344, 1246752, https://doi.org/10.1126/science.1246752, 2014.
Purvis, A., Gittleman, J., Cowlishaw, G., and Mace, G.: Predicting extinction risk in declining species, Proceedings of the Royal Society B: Biological Sciences, 267, 1947–1952, https://doi.org/10.1098/rspb.2000.1234, 2000.
Reddy, S. and Dávalos, L.: Geographical sampling bias and its implications for conservation priorities in Africa, Journal of Biogeography, 30, 1719–1727, https://doi.org/10.1046/j.1365-2699.2003.00946.x, 2003.
Tanalgo, K. C.: Open and FAIR data sharing are building blocks to bolster biodiversity conservation in Southeast Asia, Biological Conservation, 307, 111192, https://doi.org/10.1016/j.biocon.2025.111192, 2025.
United Nations Environment Programme (UNEP): COP15 ends with landmark biodiversity agreement, https://www.unep.org/news-and-stories/story/cop15-ends-landmark-biodiversity-agreement (last access: 26 January 2026), 2022.
Veach, V., Di Minin, E., Pouzols, F., and Moilanen, A.: Species richness as criterion for global conservation area placement leads to large losses in coverage of biodiversity, Diversity and Distributions, 23, 715–726, https://doi.org/10.1111/ddi.12571, 2017.
Wilson, D. and Mittermeier, R.: Handbook of the Mammals of the World, Vol. 1: Carnivores, Lynx Edicions, Barcelona, ISBN 978-84-96553-49-1, 2009.
Wilson, D. and Mittermeier, R.: Handbook of the Mammals of the World, Vol. 2: Hoofed Mammals, Lynx Edicions, Barcelona, ISBN 978-84-96553-77-4, 2011.
Wilson, D. and Mittermeier, R.: Handbook of the Mammals of the World, Vol. 4: Sea Mammals, Lynx Edicions, Barcelona, https://doi.org/10.1093/jmammal/gyv071, 2014.
Wilson, D. and Mittermeier, R.: Handbook of the Mammals of the World, Vol. 5: Monotremes and Marsupials, Lynx Edicions, Barcelona, https://doi.org/10.1093/jmammal/gyw012, 2015.
Wilson, D. and Mittermeier, R.: Handbook of the Mammals of the World, Vol. 8: Insectivores, Sloths and Colugos, Lynx Edicions, Barcelona, ISBN 978-84-16728-08-4, 2018.
Wilson, D. and Mittermeier, R.: Handbook of the Mammals of the World, Vol. 9: Bats, Lynx Edicions, Barcelona, ISBN 978-84-16728-19-4, 2019.
Wilson, D., Lacher, T., and Mittermeier, R.: Handbook of the Mammals of the World, Vol. 6: Lagomorphs and Rodents, Lynx Edicions, Barcelona, ISBN 978-84-941892-3-4, 2016.
Wilson, D., Lacher, T., and Mittermeier, R.: Handbook of the Mammals of the World, Vol. 7: Rodents II, Lynx Edicions, Barcelona, ISBN 978-84-16728-04-6, 2017.
WWF: Living Planet Report 2024, WWF, Gland, Switzerland, https://livingplanet.panda.org (last access: 26 January 2026), 2024.
Yesson, C., Brewer, P. W., Sutton, T., Caithness, N., Pahwa, J. S., Burgess, M., Gray, W. A., White, R. J., Jones, A. C., and Bisby, F. A.: How global is the Global Biodiversity Information Facility?, PLoS ONE, 2, e1124, https://doi.org/10.1371/journal.pone.0001124, 2007.
- Abstract
- Copyright statement
- Introduction
- Data and methods
- Results
- Priority-setting applications
- Discussion
- Code and data availability
- Concluding Remarks
- Appendix A
- Appendix B: Flowchart of the main processing algorithm
- Author contributions
- Competing interests
- Disclaimer
- Acknowledgements
- Financial support
- Review statement
- References
- Abstract
- Copyright statement
- Introduction
- Data and methods
- Results
- Priority-setting applications
- Discussion
- Code and data availability
- Concluding Remarks
- Appendix A
- Appendix B: Flowchart of the main processing algorithm
- Author contributions
- Competing interests
- Disclaimer
- Acknowledgements
- Financial support
- Review statement
- References